GFP treasure hunt introduction
Introduction
What does a biological sequence look like and what can you do with it?
Imagine that the EMBL Genomics Core Facility has sequenced DNA samples of different fluorescent proteins. Before starting your bioinformatics analysis on these samples, have a read below to find out how results from DNA sequencing look like.
DNA sequencing results can be visualised in two different ways:
1. Sequencing chromatograms
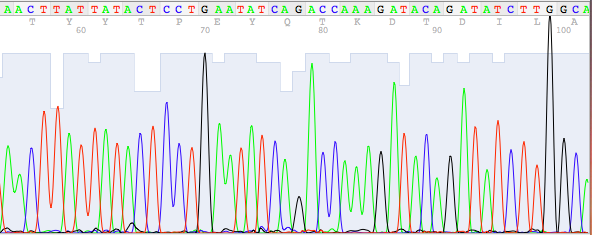
A sequencing chromatogram is displaying the data produced by the sequencing machine as so-called “peaks”.
A different colour is assigned to each of the 4 bases:
A = green , C = blue , G = black, T = red
The DNA sequence can be “read” as a sequence of individual peaks. The following is an example of such a sequencing chromatogram:
2. Text files in FASTA format
The FASTA format is a text-based format to display and store nucleic acid sequences (DNA and RNA) and amino acid sequences (protein). Each sequence starts with a single line containing a greater-than sign (>) and a descriptive header (this is the name of the sequence). The name of the sequence is added without introducing a space. The following lines contain the sequence data. The following shows an example of a sequence in FASTA format:
>Human_1233_P1 AGGAGCAGGGAGGGCAGGAGCCAGGGCTGGGCATAAAAGTCAGGGCAGAGCCATCTATTGCTTACATTTGCTTCTGACACAACTGTGTTCACTAGCAACCTCAAACAGACACCATGGTGCACCTGACTCCTGAGGAGAAGTCTGCCGTTACTGCCCTGTGGGGCAAGGTGAACGTGGATGGAAGTTGGTGGTGAGGCCCTGGGCAGGTTGGTATCAAGGTTACAAGACAGGTTTAAGGAGACCAATAGAAACTGGGCATGTGGAGACAGAGAAGACTCTTGGGTTTCTGATAGGCACTGACTCTCTCTGCCTATTGGTCTATTTTCCCACCCTTAGGCTGCTGGTGGTCTACCCTTGGACCCAGAGGTTCTTTGAGTCCTTTGGGGATCTGTCCACTCCTGATGCTGTTATGGGCAACCCTAAGGTGAAGGCTCATGGCAAGAAAGTGCTCGGTGCCTTTAGTGATGGCCTGGCTCACCTGGACAACCTCAAGGGCACCTTTGCCACACTGAGTGAGCTGCACTGTGACAAGCTGCACGTGGATCCTGAGAACTTCAGGGT
Within FASTA sequences 1-letter codes are used for nucleic acids and amino acids:
1-letter codes for nucleic acids:
Adenine = A
Cytosine = C
Guanine = G
Thymine = T
1-letter codes for amino acids:
Alanine A, Arginine R, Asparagine N, Aspartate D, Cysteine C, Glutamine Q, Glutamic acid E, Glycine G, Histidine H, Isoleucine I, Leucine L, Lysine K, Methionine M, Phenylalanine F, Proline P, Serine S, Threonine T, Tryptophane W, Tyrosine Y, Valine V
Activity navigation
Share:
 Čeština
Čeština Français
Français Ελληνικά
Ελληνικά Italiano
Italiano Svenska
Svenska