E. coli
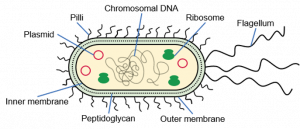
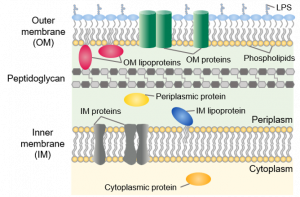
If you decide to express a recombinant protein in E. coli, the first step will be to design your expression construct and choose an expression vector. In E. coli, you can express your protein of interest in the cytoplasm or in the periplasm.
In E. coli, the cytoplasm is a reducing environment, whereas the periplasm is an oxidizing environment that allows the formation of disulfide bonds. The periplasm also contains the DsbA/DsbB oxido-reductases and the DsbC/DsbD disulfide bond isomerases. Therefore, the periplasm might be a more suitable compartment for the expression of disulfide bond-rich proteins in E. coli. Other advantages are the lower proteolytic activity in the periplasm and the presence of molecular chaperones such as FkpA, SurA and Skp. The periplasm only contains about 4% of the total cellular proteins and hence facilitates purification after an osmotic shock. The disadvantages are that protein expression in the periplasm often results in a lower yield than cytosolic expression and usually not all expressed protein will be secreted into the periplasm. To direct a recombinant protein to the periplasm, you need to add a periplasmic signal sequence to the N-terminus of your protein, which will be removed after crossing the inner membrane.
If you want to express multiple proteins simultaneously in E. coli, you can either go for monocistronic or polycistronic co-expression. Monocistronic means that each mRNA will encode for 1 protein, while in polycistronic co-expression 1 mRNA will encode for 2 or more proteins.
Co-expression can be useful if your protein of interest is part of a multi-subunit complex and might not be soluble on its own. Sometimes co-expression with chaperones or foldases can also help with in vivo protein folding. Some well-characterized chaperones are for example GroEL-GroES, DnaK-DnaJ-GrpE and tig. Chaperone plasmid sets for co-expression are also commercially available. Foldases that play an important role in protein folding are peptidyl prolyl cis/trans isomerases (PPI’s), disulfide oxidoreductase (DsbA), disulfide isomerase (DsbC) and protein disulfide isomerase (PDI).
When you co-express various proteins using multiple plasmids, the plasmids should have different antibiotic resistances and compatible origins of replication. Commercial systems for protein co-expression in E. coli are for example the pET-Duet suite and the MultiColi system.
In the pET-Duet suite, each vector has 2 individual gene expression cassettes, which allows you to co-express up to 8 proteins when combining vectors.
| Vector | Ori | Antibiotic resistance | Copy number |
|---|---|---|---|
| pET-Duet | ColE1 | Ampicillin | ~40 |
| pACYC-Duet | P15A | Chloramphenicol | 10-12 |
| pCDF-Duet | CloDF13 | Streptinomycin/spectinomycin | 20-40 |
| pRSF-Duet | RSF1030 | Kanamycin | >100 |
| pCOLA-Duet | COLA | Kanamycin | 20-40 |
A big advantage of protein expression in E. coli is the availability of a large variety of E. coli expression strains, that all have their own specific characteristics. Some strains possess extra copies of rare tRNA’s, which is very useful if the codon usage of your gene of interest is very different from the E. coli codon usage. Other strains are better equipped to deal with the expression of toxic proteins or are more suitable for the expression of disulfide bond-rich proteins in the cytoplasm. Usually, you screen a number of different expression strains and expression conditions in small scale expression and purification tests. In these small scale tests you’ll assess the total expression level of your protein of interest, the solubility and the ability to enrich it on affinity beads. Once you identify the best conditions, you can scale up your expression cultures and proceed with the large scale protein purification.
References
Kaur J., Kumar A. and Kaur J. (2018) Strategies for optimization of heterologous protein expression in E.coli: roadblocks and reinforcements. Int J Biol Macromol. 106:803-822
Rosano G.L. and Ceccarelli E.A. (2014) Recombinant protein expression in Escherichia coli: advances and challenges. Front. Microbiol. 5: 172
Rosano G.L., Morales E.S. and Ceccarelli E.A. (2019) New tools for recombinant protein production in Escherichia coli: a 5-year update. Protein Science 28: 1412-1422
Berkmen M. (2012) Production of disulfide-bonded proteins in Escherichia coli. Protein Expression and Purification 82:240-251
Vincentell R. and Romier C. (2013) Expression in Escherichia coli: becoming faster and more complex. Current Opinion in Structural Biology 23:326–334
Jia B. and Jeon C.K. (2016) High-throughput recombinant protein expression in Escherichia coli: current status and future perspectives. Open Biol. 6:160-196