The two systems we currently use at EMBL PEPCF for multi-protein expression in insect cells are the MultiBac and the biGBac systems, although various other systems based on restriction/ligation cloning, ligation-independent cloning and USER cloning exist as well. For example, the MacroBac system uses a Biobricks-type assembly method compatible with either restriction/ligation cloning (MacroBac11 series) or ligation-independent cloning (MacroBac438 series). Additionally, the MacroBac vectors are available with different N- and C-terminal tags already cloned in.
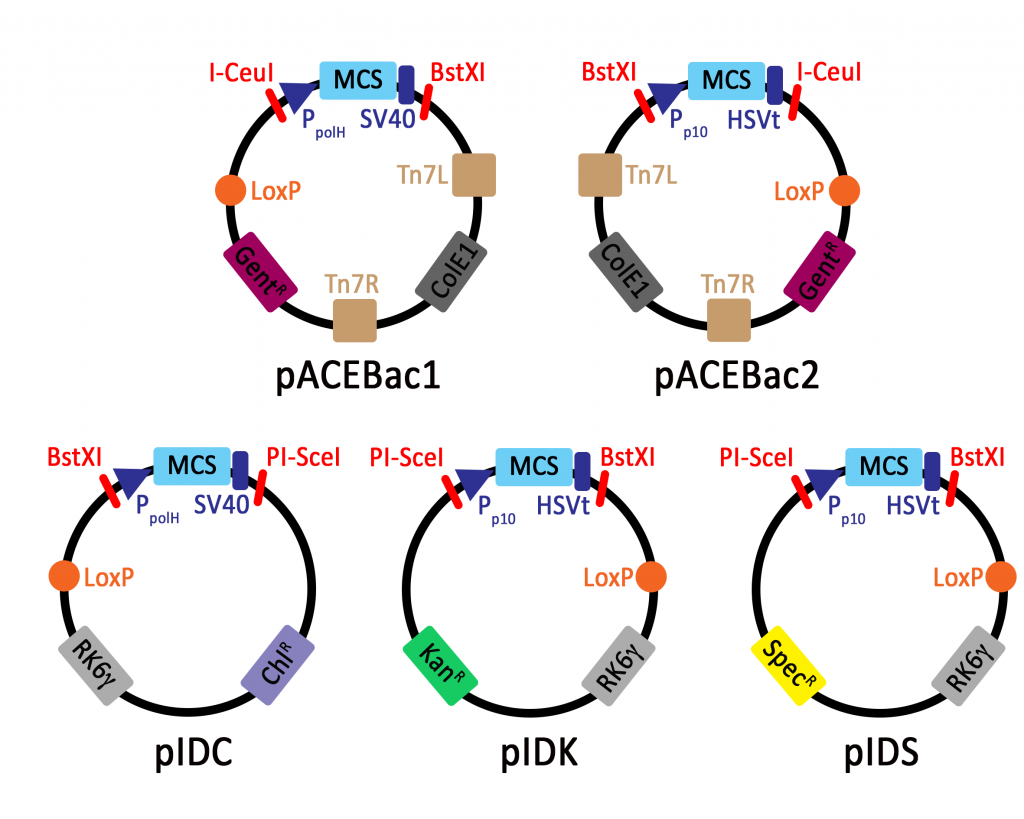
The MultiBac system has been developed by the group of Imre Berger and exists of a set of acceptor and donor vectors, that allow a flexible and modular combination of different heterologous genes. After cloning the genes of interest into the multiple cloning sites of the donor and acceptor vectors, the individual gene expression cassettes can be combined either via Cre-Lox recombination or via the multiplication modules.
The acceptor vectors pACEBac1 and pACEBac2 both contain the Tn7L and Tn7R sites for transposition into the bacmid and an I-CeuI/BstXI multiplication cassette. The donor vectors pIDC, pIDK and pIDS all contain the conditional R6Kγ origin of replication, which means they can only be propagated in pir+ bacterial cells. The multiplication cassette in the donor vectors consists of the homing endonuclease PI-SceI and BstXI restriction sites.
More information about how to use the MultiBac system can be found on the websites of Geneva Biotech and the University of Bristol.
The biGBac system uses Gibson assembly cloning for generating multi-protein expression constructs. In a first step, the genes of interest are cloned into the transfer vector pLIB. Each individual gene expression cassette (GEC) is then amplified by PCR using predefined sets of oligos that have overhangs compatible with Gibson assembly. Up to 5 individual GEC’s can be combined in a pBIG1 vector into a polygene expression cassette (PGC). The Gibson assembly reaction introduces SwaI restriction sites in between the individual GEC’s and PmeI restriction sites flanking the newly generated PGC. For protein complexes larger than 5 subunits, various PGC’s can be created into the pBIG1a-e vectors. The PGC’s excised by PmeI from the pBIG1 vectors can then be further combined with a Gibson assembly reaction into the pBIG2 vectors. Again, up to 5 PGC’s can be combined together, allowing the assembly of constructs encoding 25 different subunits. The resulting pBIG2 vectors can be analyzed via PacI (and/or SwaI) restriction digest followed by agarose gel electrophoresis.
References
Berger I., Tölzer C. and Gupta K. (2019) The MultiBac system: a perspective. Emerg Top Life Sci. 3(5):477-482
Gupta K., Tölzer C., Sari-Ak D., Fitzgerald D.J., Schaffitzel C. and Berger I. (2019)MultiBac: Baculovirus-Mediated Multigene DNA Cargo Delivery in Insect and Mammalian Cells. Viruses. 11(3):198
Sari D., Gupta K., Thimiri Govinda Raj D.B., Aubert A., Drncová P., Garzoni F., Fitzgerald D. and Berger I. (2016) The MultiBac Baculovirus/Insect Cell Expression Vector System for Producing Complex Protein Biologics. Adv Exp Med Biol. 896:199-215
Bieniossek C., Imasaki T., Takagi Y. and Berger I. (2012) MultiBac: expanding the research toolbox for multiprotein complexes. Trends Biochem Sci. 37(2):49-57
Weissmann F., Petzold G., VanderLinden R., Huis in ‘t Veld P.J., Brown N.G., Lampert F., Westermann S., Stark H., Schulman B.A. and Peters J.-M. (2016) biGBac enables rapid gene assembly for the expression of large multisubunit protein complexes. PNAS. 113(19): E2564-2569
Weissmann F. and Peters J.-M. (2018) Expressing Multi-subunit Complexes Using biGBac. Methods Mol Biol. 1764:329-343
Gradia S.D., Ishida J.P., Tsai M.-S., Jeans C., Tainer J.A. and Fuss J.O. (2017) MacroBac: New Technologies for Robust and Efficient Large-Scale Production of Recombinant Multiprotein Complexes. Methods Enzymol. 592:1-26