To determine the optimal expression conditions for your protein of interest, you will most likely start with small scale expression and purification tests to assess the total expression level of your protein of interest, the solubility and the ability to enrich your protein on affinity resin. This small scale screening can either be performed in plate format (e.g. in 24-well plates) or in small culturing flasks.
The following parameters can be varied in small scale screening experiments:
- E. coli expression strain
- E. coli growth medium
- Growth/expression temperature
- Cell density (OD600) at the time of induction
- Amount of inducer
- Amount of time in which protein expression takes place (e.g. 2 hours, 4 hours, overnight etc.)
The results of the screening experiment can be analysed via SDS-PAGE. The minimal samples you should analyse for each condition are the total lysate, the soluble fraction (after cell lysis and centrifugation) and the elution fraction from the affinity resin.
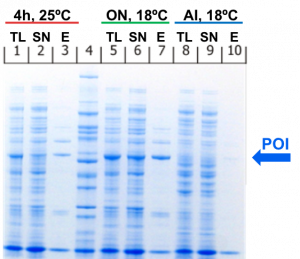
The SDS-PAGE at the right shows an example of what the results of a small scale screening could look like. In this case, a His6-tagged protein was expressed in E. coli BL21(DE3) codon + RIL in 3 different expression conditions. After cell lysis and centrifugation, a small scale affinity purification was performed with Ni-NTA beads.
Condition 1:
- Cell culture medium: LB
- Culturing parameters: grow at 37ºC until OD600 ~ 0.6 → induce with 0.5 mM IPTG → express for 4h at 25ºC
Condition 2:
- Cell culture medium: LB
- Culturing parameters: grow at 37ºC until OD600 ~ 0.6 → induce with 0.2 mM IPTG → express overnight at 18ºC
Condition 3:
- Cell culture medium: TB-FB + 0.05% glucose + 1.5% lactose + 2 mM MgSO4 (auto-induction medium)
- Culturing parameters: grow at 37ºC until OD600 ~ 0.8 → reduce temperature to 18ºC for overnight expression
In this case, the second condition obviously leads to the highest amount of soluble protein and would therefore be used for the large scale expression and purification.
At the EMBL Protein Expression and Purification Facility, we have an automated liquid handling system (Biomek NXP Span8 from Beckman Coulter) that has been adapted to perform high-throughput small scale purifications using RoboColumns, which can be packed with various resin materials. This allows us to do up to 96 small scale purifications in parallel.
Troubleshooting
Unfortunately, not all proteins you attempt to express will be soluble. When confronted with insoluble proteins, there are a number of factors you can try to change to improve solubility.
In first instance, you can try to change the expression conditions. Expression at lower temperatures or a reduction in the amount of inducing reagent might increase the amount of soluble protein you obtain. You can also try co-expression with chaperones/foldases or other expression strains with different characteristics that might be more suitable for your protein of interest.
If varying the expression conditions and/or the expression strain doesn’t help, you might consider redesigning your construct by for example including solubility-enhancing tags or using a plasmid backbone with a weaker promoter (e.g. trc or tac promoter instead of T7 promoter). Finally, changing the host organism used for protein expression might also improve the solubility of your protein. Often insect cells and mammalian cells are more suitable for expressing complex eukaryotic proteins than bacteria.