
New insights into RNA Polymerase I
Cryo EM reconstruction of RNA Polymerase I reveals details of how molecule binds and transcribes DNA
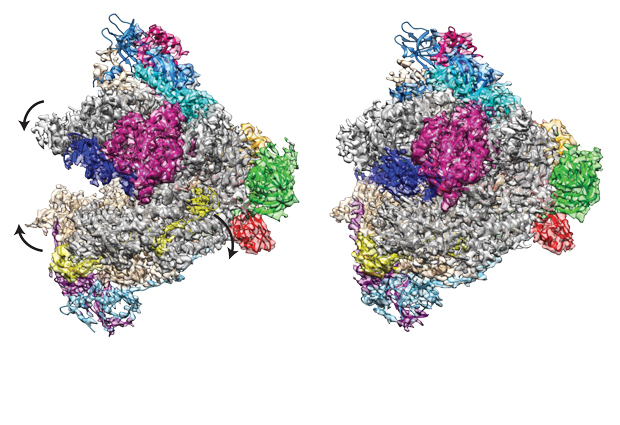
Research published today in the journal Molecular Cell sheds light on the workings of RNA Polymerase I (Pol I). Using cryo-electron microscopy (cryo EM) techniques, scientists from the Müller and Sachse groups at EMBL have now resolved the structure of Pol I to reveal details of just how the molecule binds to and transcribes DNA to produce stretches of RNA.
Pol I is one of three RNA polymerases found in many organisms and all three are central to the workings of the cell. Pol I produces stretches of RNA that together with other proteins go on to form the ribosome, the cell’s protein factory. As protein production is crucial for cells to grow and proliferate, faulty regulation of Pol I has been linked to different types of cancer.
Scientists knew the overall structure of Pol I, thanks to data from crystallography experiments. But no-one had yet managed to see how it interacts with DNA and RNA. The paper published today includes insights into the different phases of binding and transcription of DNA to RNA involved in the Pol I transcription cycle and make it possible to compare it with similar structures already known from other polymerases.


