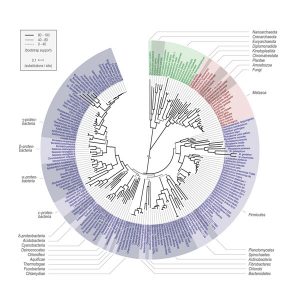
A new tree of life allows a closer look at the origin of species
A global evolutionary map reveals new insights into our last common ancestor
In 1870 the German scientist Ernst Haeckel mapped the evolutionary relationships of plants and animals in the first ‘tree of life’. Since then scientists have continuously redrawn and expanded the tree adding microorganisms and using modern molecular data, yet, many parts of the tree have remained unclear. Now a group at the European Molecular Biology Laboratory (EMBL) in Heidelberg has developed a computational method that resolves many of the open questions and produced what is likely the most accurate tree ever. The study, which appears in the current issue of the journal Science, gives some intriguing insights into the origins of bacteria and the last common universal ancestor of all life on earth today.
“DNA sequences of complete genomes provide us with a direct record of evolution”, says Peer Bork, Associate Coordinator for Structural and Computational Biology at EMBL, whose group carried out the project. “For a long time the overwhelming amount of data (the human genome alone contains enough information to fill 200 telephone books) has made it very difficult to pinpoint the information needed for a high-resolution map of evolution. But our study shows how this challenge can be tackled by combining different computational methods in an automated process.”
Bork’s lab specialises in the computational analysis of genomes, and recently they applied this expertise to the tree of life. Since all organisms descend from the same ancestor, they share some common genes. Francesca Ciccarelli and Tobias Doerks of Bork’s group managed to identify 31 genes with clear relatives in 191 organisms, ranging from bacteria to humans, to reconstruct their relationships.
“Even using such genes, you might get the wrong answer,” says Ciccarelli. “Organisms inherit most genes from their parents, but over the course of evolution, a few have been obtained when organisms swapped genes with their neighbours in a process called horizontal gene transfer (HGT). Obviously, the latter class of genes does not tell you anything about your ancestors. The trick was to identify and exclude them from the analysis.”
“This procedure drastically reduced the ‘noise’ in the data, making it possible to identify as yet unknown details of early evolution,” says Tobias Doerks. “For example, we now know that the first bacterium was probably a type called gram-positive and likely lived at high temperatures – suggesting that all life arose in hot environments.”
The improved tree has also shed light on other research carried out by the group. Bork and colleagues are participating in projects that collect genetic material of unknown species en masse from environments such as farm soil and ocean floor. “With the new high-resolution tree in hand, it is now possible to classify genetic material from this unexplored microbial world and further our understanding of life on the planet.”