
A tool to improve cancer research
EMBL scientists use ‘mini organs’ to clarify how cells turn cancerous; insights may shed light on ways to stop tumour growth
If we can spot a tumour right when it starts and know its next move, maybe we can also stop it from growing at all.
EMBL scientists have created a new, realistic 3D testbed that could help them achieve that goal by studying cancer cells as they first form. Their results are published today in eLife.
By creating ‘mini organs’, or organoids, from healthy mouse breast tissue and employing laser light-sheet technology to steadily monitor cell growth, the researchers were able to track, study, and even determine a tipping point when cells that express oncogenes (cancer-causing genes) turn cancerous. The hope is that this work could lead to new strategies for approaching breast cancer treatment.
“Organoids and viruses have been used before, but this is the first time we were able to ‘multiplex’ technologies to be able to turn on and off oncogenes and image them in real time for so long,” says Ashna Alladin, lead author on the paper and a postdoc in EMBL’s Jechlinger group at the time of the study. “This research doesn’t offer a new treatment option, but it does open the door to test new ideas in a realistic platform and get a more accurate look at tumour cell behaviour.”
While many cancer studies look at a stage of already established tumours consisting of mutated cells – those that have changes in their DNA – this research targeted the very first stages of disease when healthy cells can influence nearby unhealthy ones. Researchers have long hypothesised that even a single modified (damaged) cell could be the start of a tumour. The researchers found, however, that one damaged cell is unlikely to overtake an otherwise healthy organ, but if it’s amongst other damaged cells, they’re more likely to multiply and form a tumour.
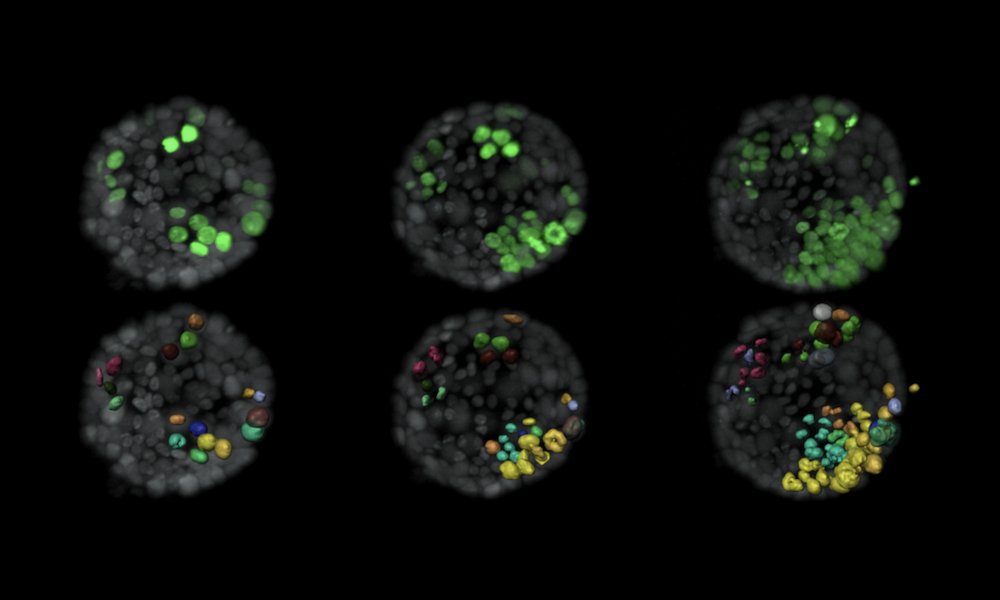
The research team grew the organoids from mouse breast duct tissue. They methodically targeted single cells by switching on oncogenes known to cause cancer in humans. They could track their work by tagging the cells with fluorescent proteins, which they illuminated with lasers, allowing them to capture snapshots every 10 minutes to study the changes at this very small scale. They adapted a type of microscope technology called selective plane illumination microscopy (SPIM) to follow these modified cells within 3D organoids for up to 3 days. SPIM uses a laser to illuminate just a single plane of a sample at a time, scanning it over the whole sample to build up a 3D image. This minimises damage to the sample caused by light, making SPIM an ideal technique for studying biological specimens over extended periods of time.
The researchers developed a computer algorithm from the stream of cell images and used it to identify patterns of behaviour in the cells that triggered tumours. One feature seemed consistently linked with tumour growth: as the ratio of modified cells to healthy cells increased – especially when modified cells were near one another – they became more likely to form tumours. The team theorised that healthy cells send signals that suppress tumour formation to modified cells. However, this suppression only works until that signal is drowned out by larger numbers of modified cells communicating among themselves.
“We’ve now created a robust platform to explore cell deficiencies and a way to realistically study response to treatments at the cellular level,” says EMBL group leader Martin Jechlinger. “If we know about deficiencies already tested in these systems, we can potentially more accurately flag risk groups for better medical care, in much the same way that identifying genes like BRCA1 and BRCA2 – which are found mutated in breast cancer – has informed medical decisions. Where this could potentially lead is quite exciting.”


