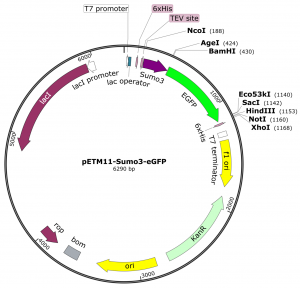
We have a modified pETM11 vector for the expression of SUMO3 fusion proteins available. This vector contains an optimized synthetic human sumo3 gene with appropiate restriction sites at the 3′-end of the sumo3 gene for the insertion of any gene of interest.
The small human SUMO3 protein (81 aa, 9.2 kDa) can be used as an N-terminal fusion partner for the expression of recombinant proteins. The SUMO3 fusion tag often enables high expression yields and good solubility of the target proteins. During the purification process, the SUMO3 fusion tag can be easily removed by a very robust and highly specific protease called SenP2, which leaves no additional amino acid overhangs on your target protein. The unique feature of this protease is the recognition of the 3-dimensional structure of SUMO3 and cleavage after the conserved Gly-Gly motif, which differs from other proteases that recognize specific amino acid sequences. We also have a vector encoding His6-tagged SenP2 (containing a codon-optimized synthetic SenP2 gene) available.
Both the pETM11-Sumo3-eGFP vector and the pETM11-SenP2 vector encoding the corresponding SUMO protease can be obtained from EMBL via the Material Transfer Agreement form.
How to clone your gene of interest into the pETM11-Sumo3-eGFP vector
The first option for inserting your target gene is by using the BamHI site at the 5’ end and any of the other downstream restriction sites at the 3’ end. In this case you will end up with an extra Serine residue at the N-terminus of your protein after SenP2 cleavage due to the TCC codon of the BamHI site. When you use the BamHI-compatible BglII site AGATCT in your insert, you will also introduce an extra Ser encoded by the TCT codon of the BglII site.
If you don’t want any additional amino acids at the N-terminus of your protein after SenP2 cleavage, you have to use the AgeI restriction site or the compatible sites for AvaI or XmaI (CCCGGG), BspEI (TCCGGA) or NgoMIV (GCCGGC).
No matter which strategy you decide to go for, you have to make sure that you have the C-terminal Gly-Gly codons of the SUMO3 gene in frame with your target gene.
Note: The first amino acid after the Gly-Gly can not be Proline, as the protease will not be able to cleave the fusion protein and remove the SUMO3 tag in that case.
Purification of the fusion protein is achieved via the N-terminal His6-tag added to the SUMO3. However, you can also obtain your target protein with a C-terminal His6-tag if you don’t add a stop codon to your gene. If you wish to include a C-terminal His6-tag, please make sure that the second His6-tag is in frame with your gene of interest. Depending on the 3′ restriction site you choose, you may have to add one additional base to remain in frame.
The pETM11-Sumo3-EGFP vector also contains an eGFP gene downstream of the SUMO3 coding sequence, which results in the following advantages:
- easy confirmation of double digest of the vector due to the release of the eGFP fragment, which is replaced subsequently by your target gene
- optional N-terminal tagging of your target protein with eGFP by using the SacI restriction site at the 5’ end of your gene of interest and one of the restriction sites downstream of the eGFP gene at the 3’ end (optional with or without C-terminal His6-tag).
- optional C-terminal tagging of your target protein with eGFP by using the AgeI and BamHI restrictions sites at the 5’ and 3’ ends of your gene of interest, respectively
References
Marblestone J.G., Edavettal S.C., Lim Y., Zuo X. and Butt T.R. (2006) Comparison of SUMO fusion technology with traditional gene fusion systems: enhanced expression and solubility with SUMO. Protein Science 15(1): 182-189
Panavas T., Sanders C. and Butt T.R. (2009) SUMO Fusion Technology for Enhanced Protein Production in Prokaryotic and Eukaryotic Expression Systems. Methods Mol Biol 497: 303-317
Reverter D. and Lima C.D. (2006) Structural basis for SENP2 protease interactions with SUMO precursors and conjugated substrates. Nat Struct Mol Biol 13(12): 1060-1068