Zeroing in on heart disease
Innovative strategy pinpoints genes underlying cardiovascular disease risk
In a nutshell:
– Genome-wide association studies (GWAS) enable scientists to trace the origin of human diseases to distinct regions in the genome, but their resolution is limited
– RNA interference is a powerful technology to test, in parallel, the function of many genes
– The combination of these two approaches pinpoints what genes are the most important for the control of cholesterol levels, and thus likely to confer a risk for cardiovascular disease
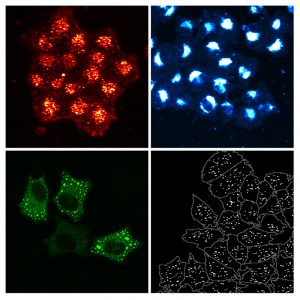
Studies screening the genome of hundreds of thousands of individuals (known as Genome-wide association studies or GWAS) have linked more than 100 regions in the genome to the risk of developing cardiovascular disease. Researchers from the European Molecular Biology Laboratory (EMBL) and the University of Heidelberg, through the joint Molecular Medicine Partnership Unit (MMPU), are taking these results one step further by pinpointing the exact genes that could have a role in the onset of the disease. Their findings are published today in the Public Library of Science (PLoS) Genetics.
The scientists used a technology called “RNA interference” that can selectively decrease the level of expression of targeted genes. By observing what changes, if any, this decrease causes in cells, researchers can identify the function of the genes and, on a larger scale, objectively test the function of many genes in parallel.
Cholesterol levels in the blood are one of the main risk factors for cardiovascular disease. They are controlled by the amount of cholesterol that cells can take in – thus removing it from the blood – and metabolise. The researchers used RNA interference to test the function of each of the genes within 56 regions previously identified by GWAS as being linked with cardiovascular disease. They selectively decreased their action and measured what, if any, changes this induced in cholesterol metabolism. From this they could deduce which of the genes are most likely to be involved in the onset of the disease.
“This is the first wide–scale RNA interference study that follows up on GWAS. It has proven its potential by narrowing down a large list of candidate genes to the few with an important function that we can now focus on in future in-depth studies,” explains Rainer Pepperkok at EMBL, who co-led the study with Heiko Runz at the University of Heidelberg.
“In principle, our approach can be applied to any disease that has an observable effect on cells”, adds Heiko Runz. “The genes identified here may further our understanding of the mechanisms leading to cardiovascular disease and allow us to improve its prediction and diagnosis”.



