
Novel method sheds new light on tumour-suppressing RNA
Researchers from EMBL Grenoble, in collaboration with colleagues at the Max Delbruck Center in Berlin and the Institut de Biologie Structurale (CEA/CNRS/UGA) in Grenoble, have developed a way to visualise large RNAs in 3D using biochemical and structural biology techniques.
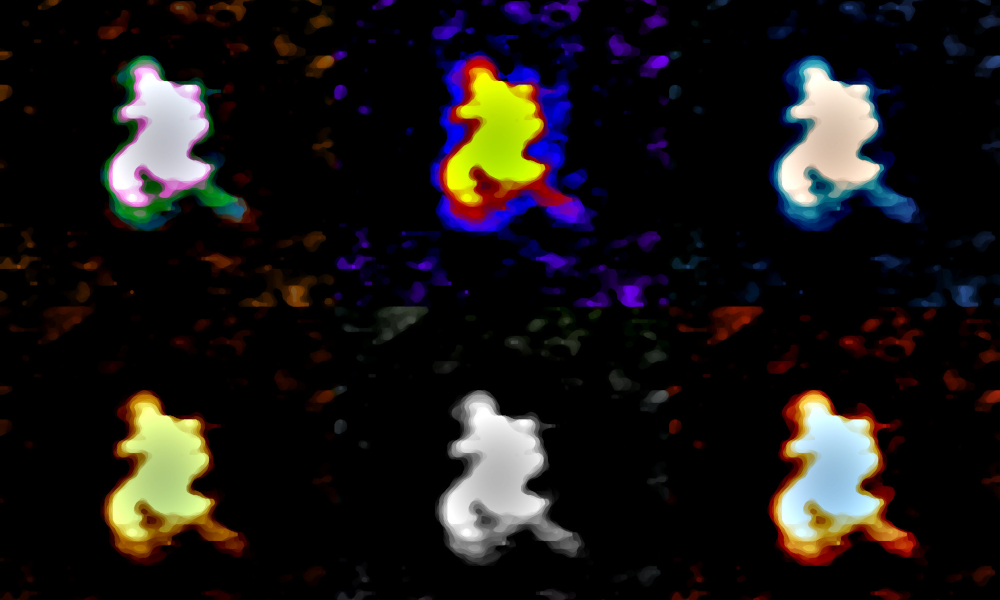
The method was used to characterise a human tumour-suppressing RNA, but may also have potential applications in viral RNA research, such as for SARS-CoV-2. The description of this new approach has been presented in Nature Protocols on 25 May 2020.
Large RNAs not only code for proteins, but also have an active role in vital cellular processes, such as cellular differentiation and DNA replication. Their proper functioning is crucial to prevent severe diseases such as cancer and neurological or cardiovascular defects. Therefore, understanding how these large RNAs work is of crucial interest for a wide range of research areas, including epigenetics, neurobiology, oncology, and infection biology.
Although arrangement of RNAs into specific three-dimensional structures is a key determinant of their biological function, the unprecedented molecular complexity of many large RNAs has so far precluded their 3D structural characterisation. The method now described by researchers at EMBL and collaborating institutions will advance research in this field.
“Our protocol describes a solid, broadly applicable method to overcome the difficulties in visualising the shapes of large RNAs. This makes their 3D characterisation accessible to a lot of labs around the world,” says Dr. Tina Uroda, lead author of the paper and former graduate student in the group of Dr. Marco Marcia at EMBL Grenoble. “This will help us get detailed insights into many medically relevant targets.”
The scientists used two different, well-established structural biology techniques: Using atomic force microscopy, they captured individual RNA particles and inferred their size and compactness; whereas they determined the three-dimensional shape of the RNA in solution by small angle X-ray scattering.
The researchers developed the protocol for a human long non-coding RNA called MEG3. However, the protocol is broadly applicable, especially considering that human cells express more than 30 000 long non-coding RNAs compared with about 20 000 RNAs encoding proteins. “When we recorded the first images of MEG3, we were stunned by its surprising compactness,” says Dr. Jean-Luc Pellequer, CEA researcher at the Institut de Biologie Structurale and co-author. “Looking at the results we obtained for MEG3, we realised that analysing long non-coding RNA images quantitatively with our protocol could reveal useful biological insights,” adds Dr. Paolo Annibale, another co-author from the Max Delbruck Center in Berlin.
Besides human RNAs, the protocol could also be applicable to structural characterisation of large viral RNAs. “RNA viruses, like the SARS-CoV-2 coronavirus that is responsible for the current COVID19 pandemic, have large untranslated regions,” explains co-author Dr. Isabel Chillón, from EMBL Grenoble. “The secondary structures of these regions – which shows multiway junctions, stem loops and pseudoknots – suggest a very complex 3D topology.” Previous studies have shown that changes in the 3D structure within these untranslated regions regulate the translation of viral structural and non-structural proteins. “This is where our work could have an important impact on this current situation,” highlights Chillón. “It would be very gratifying if our protocol now helped to characterise the SARS-CoV-2 RNAs and their conformational changes.” Such studies would shed light on key aspects of the viral life cycle and perhaps promote the development of targeted drugs in the future.


