
Breaking boundaries
Alumnus Stefan Hell jointly received this year's Nobel Prize in Chemistry, for breaking barriers in light microscopy. This pioneering discovery shapes the work of EMBL scientists – how and why, and what's next?
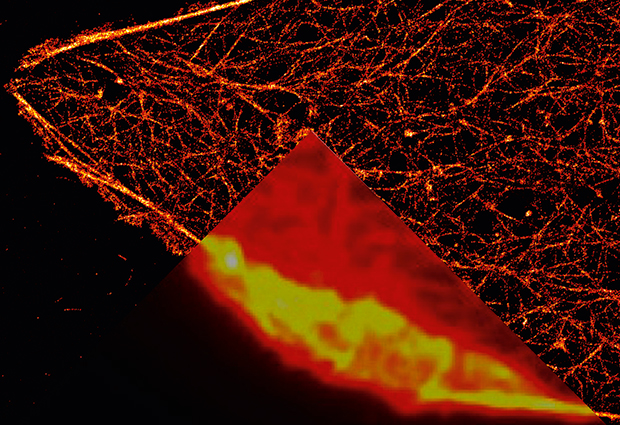
In the 1600s, the first microscopes allowed people to see a world hidden until then by the limits of the human eye: amoeba, sperm cells, the details of insects’ wings. It took almost 400 years for researchers to push the optical microscope deeper, bringing into focus individual molecules and the inner workings of cells.
The Royal Swedish Academy of Sciences awarded the 2014 Nobel Prize in Chemistry to three of the researchers responsible for inventing what is called nanoscopy or super resolution microscopy, recognising Eric Betzig, William Moerner and Stefan Hell.
The three were able to circumvent something called the Abbe limit, once considered a hard minimum on the smallest object an optical microscope could distinguish. An equation formulated in 1873 by German physicist Ernst Abbe showed it was impossible to resolve two objects closer than half of light’s wavelength, about 200-350 nanometers. That’s enough to clearly see a whole human cell or bacterium, but not a virus or a protein. While electron microscopes – developed in the 1920s and 30s – can see much smaller than that, they require samples to be set in preservative or frozen and sliced into thin layers, and therefore cannot be used to look at living cells.

Ingenious solution
Working first at the University of Turku in Finland and then at the Max Planck Institute for Biophysical Chemistry in Göttingen, Hell – who had previously done postdoctoral research at EMBL Heidelberg – was one of the first to show that this boundary on optical microscope resolution could be circumvented if dyes with special photochemical properties are used, raising the tantalising possibility of a microscope that achieved a level of detail close to that of electron microscopy without its drawbacks. He did it by limiting the area in which the dyes responded to light to a region narrower than the resolution limit, circumventing the Abbe limit.
We are able to see structures and the dynamics of structures that we have never been able to see before.
First, Hell used fluorescent dyes – compounds that glow when exposed to light – to tag specific molecules within cells. Then, he shined two lasers on the sample, one that made the dye glow in a focused point, and a second in a donut shape around that point which cancelled out fluorescence from most of the area except for a nanometre-sized central spot. By moving this effectively ‘sharpened’ laser beam across a sample, Hell could piece together clear images of structures below Abbe’s theorised limit. Betzig and Moerner achieved a similar result by selectively activating single fluorescent molecules a few at a time, then combining many such images to get a clearer, more highly resolved picture.
“We all learned in high school that you cannot use a microscope beyond half the wavelength of the light the specimen emits,” says Rainer Pepperkok, head of the EMBL Core Facilities and Advanced Light Microscopy Core Facility, and Team Leader in the Cell Biology and Biophysics Unit. “We are just now getting beyond this barrier and therefore we are able to see structures and the dynamics of structures that we have never been able to see before.”
Watching life move
Pepperkok is among many EMBL scientists who use microscopy in their work. His group studies how molecules move across the membranes of cells and between organelles. The structures that control movement between organelles are only 50 to 60 nanometers across – about 2000 times smaller than the thickness of a human hair – while other structures are within a few nanometers of each other, well below the limit of ordinary optical microscopy. “Live cell imaging is a must for us,” he explains, “because we want to study how these things move.” That’s why electron microscopy – which can pinpoint such nanostructures but only works well with very thin slices of fixed cells – is also out of the question.
His group needs to see how the molecules interact in functioning cells to truly understand what is happening. “Work in a living cell was for many years hampered because an ordinary light microscope does not provide the resolution that is required to resolve the structure that we are interested in,” he says. With super-resolution microscopy, observing live cells at that level of detail is now possible.
That capability has allowed Edward Lemke, group leader in the Structural and Computational Biology Unit, to better understand the proteins that control traffic in and out of a cell’s nucleus. Lemke explains that the proteins that regulate this process are disordered in their natural state inside the cell, meaning they don’t have a rigid three-dimensional structure that computers can predict or that can be easily determined by structural biology techniques, such as those used at EMBL Hamburg and EMBL Grenoble. In future, Lemke and his group would like to use super-resolution microscopy to see how such proteins act in the dynamic conditions inside a cell.
“The idea of moving biochemistry into the living cell is dramatically facilitated by super-resolution microscopy,” he says. By turning cells into test tubes, scientists can learn much more about the way protein interactions and other biochemical processes unfold in the real world.
Lemke’s group have also used the quantitative capabilities of super-resolution microscopy. “There were estimates that the nuclear pore protein complex had around 1000 proteins, but no one really knew exactly how many there were,” Lemke explains. From the intensity and frequency of the dye-tagged proteins’ glow, the group was able to determine exacly how many proteins the complex contained. “This was something we couldn’t have done before super-resolution microscopy, something I don’t know any other way we could have done.”
Expanding into space
Jonas Ries is another young scientist pushing the boundaries of microscopy. A group leader in the Cell Biology and Biophysics Unit, Ries is working to improve the three-dimensional capabilities of super-resolution microscopy. His group has found a way to determine a glowing molecule’s position in space. By looking at fluorescence generated by a molecule’s interaction with the glass plate behind it in addition to its normal fluorescence, they can find each molecule’s location in space, allowing them to make a three-dimensional model of the structure they’re studying.
He’s also exploring what can be gained by combining super-resolution microscopy with other imaging technology, like electron microscopy or light sheet microscopy, which excites fluorescence only in a thin slice of a sample. For instance, Ries envisions learning more about endocytosis – which is how cells engulf and ‘swallow’ molecules – by looking closely at each of the 50 or so proteins involved using electron microscopy and then determining their positions relative to one another from the more zoomed-out view super-resolution microscopy allows.

Jan Ellenberg, Head of the Cell Biology and Biophysics unit, points to Ries’ work as an example of how researchers are using super-resolution microscopy in inventive and ever more useful ways. He utilises the technique himself to study the organisation of the cell’s nucleus. Last year, his group was among the first to demonstrate super-resolution microscopy’s power for structural biology when they published how the different proteins inside nuclear pore protein complexes are arranged.
“With super-resolution microscopy you can start to see the internal molecular organisation of individual large protein complexes,” Ellenberg says. “That was a very controversial matter before: we could not directly visualise that inside a cell.”
More to come
One thing he says will make super-resolution microscopy more useful are better fluorescent dyes – his colleagues agree: “Super-resolution microscopy in principle can make clear images down to a few nanometers,” Pepperkok explains. “However, this is at a price; you need a lot of energy, and living cells cannot cope with that.” With dyes that produce better, more distinguishable signals and are more tolerable to living cells, he and other EMBL scientists say that they will soon learn more about structural biology, cellular division and more.
This is at a price; you need a lot of energy, and living cells cannot cope with that
That could mean a whole new depth of understanding in biochemistry when it’s occurring inside living matter. And, even though super-resolution microscopy is still the domain of fundamental – rather than applied – research, Pepperkok points out that work already under way by Stefan Hell’s group at the German Cancer Research Center could help answer important questions about the disease. For researchers at EMBL and around the world, super-resolution microscopy has lifted the veil further on the once-mysterious inner workings of our cells.


