
Understanding chromosome organisation
EMBL scientists and collaborators help reveal the process by which enormous quantities of DNA are folded into cells
Our cells contain vast amounts of DNA. But the process through which DNA is carefully ordered and compacted has eluded scientists for decades. Now, a team coordinated by EMBL has helped to shed light on the way in which so much DNA is able to fold in such an ordered way.
It’s hard to imagine just how much DNA gets squeezed into each of our cells. The figures tell us that about two metres of DNA are organised in a way that allows them to fit inside the nucleus of a cell measuring just 5–10 micrometres (thousandths of a millimetre) across. In fact, the process of DNA folding to fit inside the nucleus is equivalent to squeezing over 70 km of thread into a single football! And what’s more, it all has to be folded in a way that allows it to function perfectly.
When DNA is folded into chromosomes, the process has to be done in an extremely structured way. The careful folding technique ensures that chromosomes can be systematically sorted out during cell division.
Imagine packing to go on holiday. You’ve neatly laid your clothes out, ready to go in the suitcase. Some people like to be precise with their packing, while some like to throw everything in and sit on the case to force it closed. With the latter approach, the clothes end up squashed together and harder to find. It might help you to get away on holiday, but it’s not an approach that cells can use to arrange their DNA.
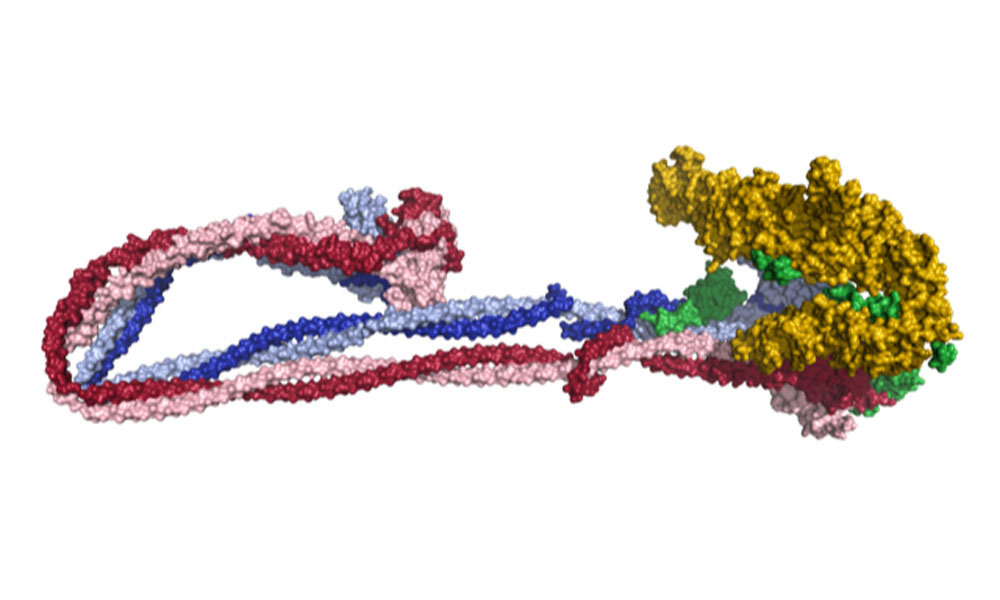
The master organiser behind this process is a protein complex called condensin. Scientists think that condensin achieves its task by looping DNA into a more compact arrangement, but the underlying molecular mechanism has remained unknown until now. A team of scientists from the Häring and Beck groups at EMBL Heidelberg, together with members of the Löwe group at the MRC Laboratory of Molecular Biology in Cambridge, UK, and the Aragon group at the MRC London Institute of Medical Sciences, have now produced the first full atomic model of condensin.
To determine the structure, they used cryo-electron microscopy (cryo-EM) – a technique pioneered at EMBL that involves rapidly freezing samples at very low temperature and imaging them using a beam of electrons.
“Gaining the first cryo-EM structures of condensin in its entirety is an amazing achievement by Fabian Merkel and Byung-Gil Lee, the two lead authors of the study, and for all colleagues involved,” says EMBL group leader Christian Häring. The work opens new opportunities for understanding the molecular movements that the condensin complex undergoes, and how its action ensures that cells partition their genetic material correctly during cell division. Häring sees a number of questions arising from finally being able to model this complex. “We next need to find out how the large conformational changes in the structure of the condensin complex are coupled to DNA looping, if we want to solve this central puzzle of genome biology.”


