
New software makes 3D imaging accessible
Open-source software allows standard microscopes to accurately image 3D structures
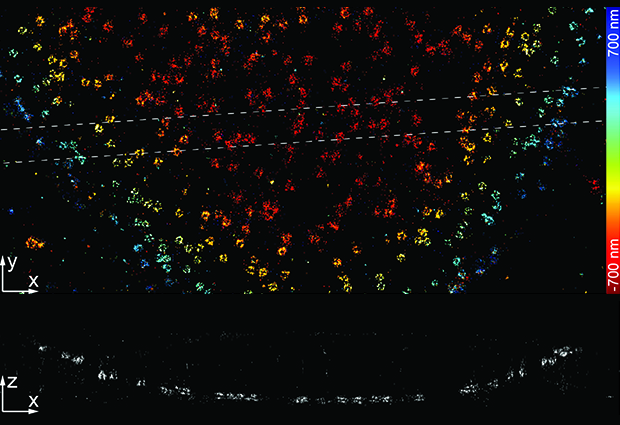
Structures inside cells, such as the nucleus and its proteins, are three dimensional. Yet scientists have often had to study such structures in 2D, because appropriately equipping microscopes was technically challenging and expensive. Jonas Ries, alongside his team and collaborators, have now published a paper in Nature Methods, showcasing their open-source 3D imaging software. This technology is used alongside standard microscopes, to capture high-quality images of 3D structures and reconstruct them in real-time, allowing more scientists to access robust 3D imaging techniques.


