
Loops, loops and loops: how DNA gets organised
Researchers from Delft University and EMBL reveal the formation of neat DNA loops by condensin protein complexes
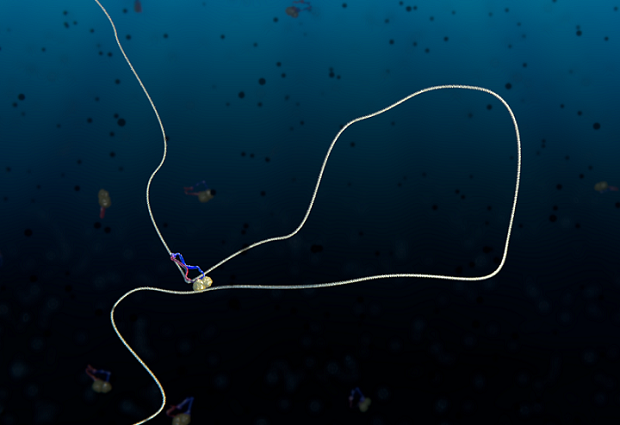
Living cells neatly package a big jumble of DNA, over two meters in length, into tidy, tiny chromosomes while preparing for cell division. Scientists have puzzled for decades over how this works. Researchers from Delft University and EMBL have finally managed to isolate and film the process, and witnessed – in real time – how a single protein complex called condensin reels in DNA to extrude a loop. By extruding many such loops in long strands of DNA, the cell packs its genome so it can be distributed evenly between its two daughter cells. The scientists published their findings online in Science on 22 February.
Spaghetti
This major discovery finally resolves a question that has been discussed in biology for over a century: how does the cell organise its spaghetti jumble of DNA into neat chromosomes in order be able to distribute its DNA evenly between two daughter cells? For many years, it has been clear that a protein complex called ‘condensin’ plays a key role, but until now, biologists were divided on exactly how condensin functioned. One theory stated that condensin works like a hook that can grasp and connect DNA within the jumble of DNA, thus tying it together. Another theory suggested that the ring-shaped condensin pulls the DNA inwards to create a loop.
Scientists from the Cees Dekker group at the Kavli Institute of Delft University, together with scientists from the group of Christian Haering at EMBL Heidelberg who established the purification and fluorescence labelling of the protein, have now managed to catch the condensin complex in the act of extruding a loop of DNA.
“This settles the debate,” says Cees Dekker. “These data provide compelling evidence that condensin indeed reels in DNA to form loops. Our novel imaging approach also allows measurement of all kinds of quantitative data: the symmetry of the loop extrusion, the speed at which the loop is formed, what happens when you pull on the DNA.”
“What is so amazing about this loop extrusion activity is the high speed by which condensin can reel in DNA, up to 1500 base pairs per second,” adds Christian Haering. “At the same time, the complex hydrolyses only a small amount of ATP”, which is the ‘fuel’ of the condensin motor – indicating that condensin does not step along the DNA base by base, but pulls it in large steps.”
Medical relevance
The research represents a significant step in the fundamental understanding of DNA in our cells, but it is also relevant to medical research. Problems with the protein family to which condensin belongs—the SMC proteins—are related to hereditary conditions such as Cornelia de Lange Syndrome. As condensin is crucial in the organisation of the chromosomes during cell division, errors in the process can result in cancer. A better understanding of these fundamental processes is vital for tracking down the molecular origins of serious illnesses.
A longer version of this story was originally published by Delft University


