Biology in four dimensions
Most things that happen in the cell are the work of ‘molecular machines’ – complexes of proteins that carry out important cellular functions. Until now, scientists didn’t have a clear idea of when proteins form these machines – are these complexes pre-fabricated or put together on the spot for each specific job? Researchers at the European Molecular Biology Laboratory (EMBL), working closely with scientists from the Technical University of Denmark (DTU), have now answered that question by drawing together many types of data in a fascinating new model. The work is published in this week’s edition of Science.
“Past studies of this type have usually left out a crucial element: time,” says EMBL Group Leader Peer Bork. “But now a picture has emerged which is extremely dynamic.”
The researchers discovered that in yeast, key components needed to create a machine are produced ahead of time, and kept in stock. When a new machine is needed, a few crucial last pieces are synthesized and then the apparatus is assembled. Holding off on the last components enables the cell to prevent building machines at the wrong times. That’s a different scenario from what happens in bacteria, which usually start production of all the parts, from scratch, whenever they want to get something done.
“We saw a clear pattern as to how the complexes are assembled,” says Sören Brunak from DTU. “It’s unusual to find such concrete patterns in biology, compared to physics for example, due to the evolutionary forces that change living systems. But using this new model, the underlying principle became very clear.
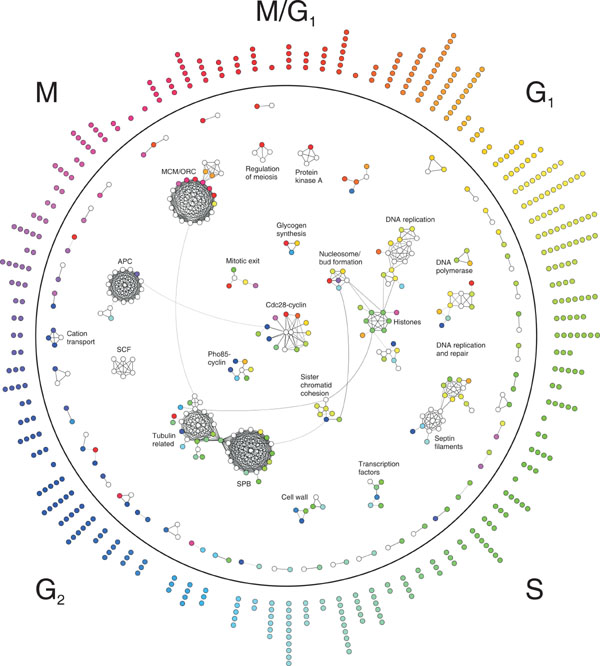
The researchers developed the model by combining data from many different studies. First, they analyzed existing protein-protein interaction data to construct a network that traces the pieces of each complex and shows how they interact. Then, to add the dimension of time, they identified genes involved in the cell cycle to find out when certain genes are switched on to produce the proteins that assemble into cellular machines. By overlaying various data sets, the researchers were able to construct a new model for protein-complex interactions.
“In addition to gaining new information about known cellular machines,” Bork notes, “we were able to plug in some components whose functions had been unknown. And the same approach could be used to study the dynamic behaviour of other biological systems, including those of humans and animals.”



