
True Love
Key proteins involved in gene expression depend on each other when forming a heart in an embryo, but, if left alone, they start to run amok.
By Dana Smith
Scientists at the Gladstone Institutes and EMBL Heidelberg have discovered that three transcription factors — proteins that turn genes on or off — interact with each other and the genome to influence how a heart forms in an embryo. Without these protein interactions, severe congenital heart defects can occur. By understanding how the transcription factors work together during heart development, researchers may discover new ways to treat heart disease.
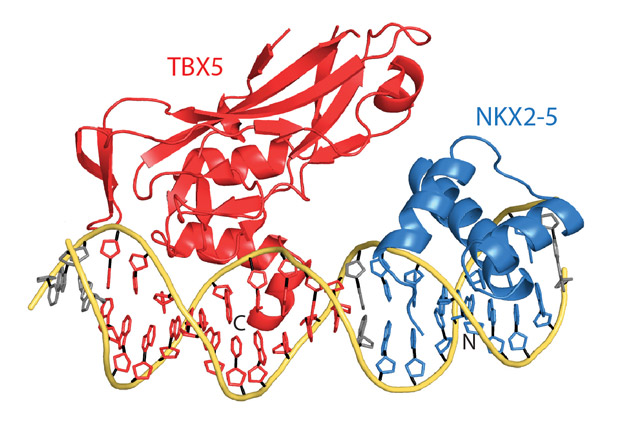
In the study, published today in Cell, Christoph Müller’s group at EMBL Heidelberg determined the 3D structure of transcription factors TBX5 and NKX2-5, bound to DNA. “The crystal structure critically shows the interaction between the two transcription factors and how they influence one another’s binding to a specific stretch of DNA,” says Müller. “In some other parts of the genome, we’re still not sure how this would work – but it would be fascinating to find out!”
For more information about the effects of these particular interactions on heart development, read the Gladstone Institutes’ press release.


