Decaying RNA molecules tell a story
Once messenger RNA (mRNA) has done its job – conveying the information to produce the proteins necessary for a cell to function – it is no longer required and is degraded. Scientists have long thought that the decay started after translation was complete and that decaying RNA molecules provided little biological information.
Now a team from EMBL Heidelberg and Stanford University led by Lars Steinmetz has turned this on its head in an article published in Cell. The researchers have shown that one end of the mRNA begins to decay while the other is still serving as a template for protein production. Thus, studying the decaying mRNA also provides a snapshot of how proteins are produced.
The discovery was made almost by accident. As part of research into how DNA is transcribed into mRNA, co-researchers Vicent Pelechano and Wu Wei developed an in vivo method to count decaying mRNA molecules in the cell. mRNA has a protective ‘cap’ that prevents it from being degraded – once that cap is removed, the decay begins. The researchers spotted a pattern in the distribution of the ‘cap-less’ RNA that they didn’t expect.

Vicent Pelechano, from the Steinmetz group at EMBL Heidelberg, explains: “The decaying RNA was thought to be of little interest biologically, so we were really surprised to see a pattern. We looked more deeply into it because it appeared to be linked to the genetic code, but we never expected it to lead to a completely new understanding of how mRNA and ribosomes interact.”
We never expected it to lead to a completely new understanding of how mRNA and ribosomes interact.
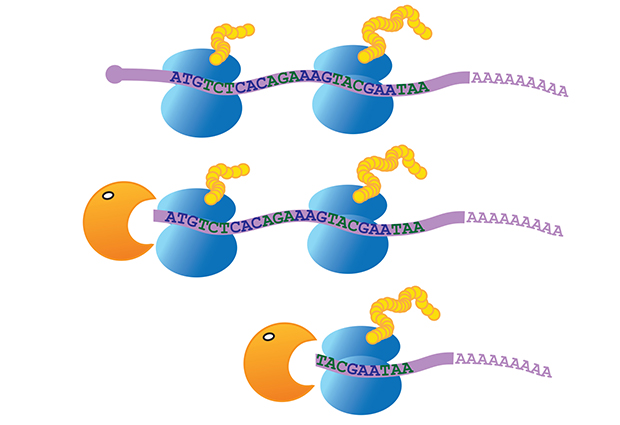
Proteins are produced from mRNA by ribosomes – ‘molecular machines’ that pass successively along the mRNA to translate its nucleotides into amino acids. It was thought that the mRNA only started to decay once the final ribosome had left it and translation was complete. In fact, the researchers were able to determine that the protective ‘cap’ is removed and degradation begins even while ribosomes are still associated with the mRNA.
They demonstrated that the enzyme degrading mRNA follows closely behind the ribosome, pausing at set points along the mRNA, usually after each group of three nucleotides, the section of code that relates to one amino acid. The team believes that this shows the enzyme pausing while translation goes on – allowing the ribosome to do its work and move on – before starting to degrade the mRNA previously protected by the ribosome.
This novel approach also opens up new avenues to study ribosomes. “Researchers studying ribosome activity usually use drugs to stall the ribosomes in place. This can alter the way we perceive their activity. We are simply looking for the molecules remaining from mRNA decay and determining their distribution. We believe this shows a more accurate picture of what is happening to the mRNA and the ribosome than was previously possible,” explains Wu Wei of Stanford University.



