Plant DNA Barcoding Resource
Overview
DNA barcoding allows the identification of living organisms on a molecular level.
This educational resource provides school students and teachers with a step-by-step guide on how to identify plant species in their own DNA barcoding project: from collecting samples to wet-lab experimentation and bioinformatics analysis. It guides barcoders through the entire process of DNA barcoding from sample collection tto wet-lab experimentation and bioinformatics analysis of sequencing data, providing detailed protocols and instructions. The resource also offers tutorials for bioinformatics analysis.
There are two alternative ways of using the resource:
1. Sample collection, wet-lab and bioinformatics analysis
This involves students collect their own biological plant material, isolate the marker gene in the lab, submit their samples for DNA sequencing and analyse the results using bioinformatics tools. These steps require access to plant material, wet-lab equipment and consumables, a Sanger sequencing service and computers with internet connection.
2. Bioinformatics analysis only
This focuses on the bioinformatics aspect of DNA barcoding without collecting samples and running wet-lab activities. The resource provides real-live example sequence data which can be used for bioinformatics analysis.
DNA Barcoding
What is DNA barcoding?
DNA barcoding is a molecular method for species identification of living organisms. The basis of this method form so-called DNA barcodes – marker genes about 300-800 nucleotides in length. Similar to industrial barcodes which identify specific products in the shop, DNA barcodes are used to discriminate between species within a group of living organisms.
Plant species identification by DNA barcoding
This educational resource offer protocols to identify plant species on a molecular level by DNA barcoding.
The two major DNA barcodes used for DNA barcoding of plants are the matK and rbcL genes; our experiment will focus on the rbcL gene.
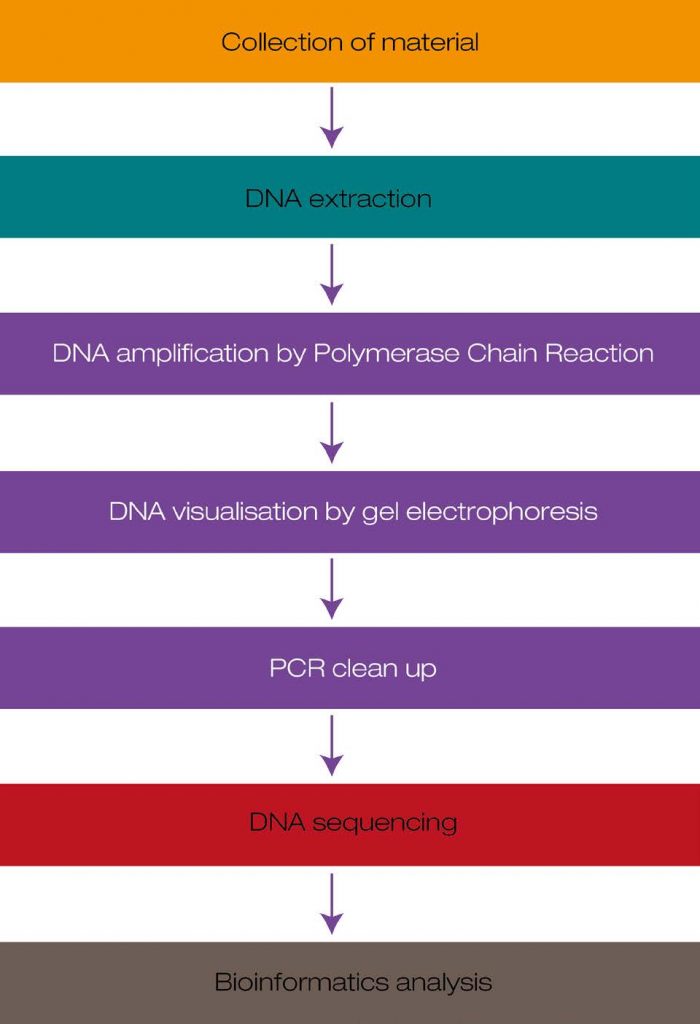
The schematic below illustrates the individual steps involved in DNA barcoding. In short, after obtaining appropriate plant material, the plant’s genomic DNA is isolated and the barcode region is amplified. Subsequently, gel electrophoresis is used to confirm the presence of the barcode. After a clean-up step to remove unwanted reagents from the PCR reaction, the PCR product is sequenced. The unique barcode sequence of the analysed plant can now be compared to barcode sequences of plant species in a nucleotide database, the European Nucleotide Archive (ENA).

Activity navigation
Sample collection and wet-lab
Bioinformatics

Topic area: Bioinformatics, Environmental biology
Age group: 16-19
Author: ELLS Team
Share: