New potential drug target in tuberculosis
Scientists reveal structure of a key tuberculosis protein
Tuberculosis remains one of the deadliest threats to public health. Every year two million people die of the disease, which is caused by the microorganism Mycobacterium tuberculosis. Roughly one third of the world’s population is infected and more and more bacterial strains have developed resistance to drugs. Researchers from the Hamburg Outstation of the European Molecular Biology Laboratory (EMBL) and the Max Planck Institute for Infection Biology (MPIIB) in Berlin have now obtained a structural image of a protein that the bacterium needs for survival in human cells. This image reveals features of the molecule that could be targeted by new antibiotic drugs.The results appear in this week’s online issue of the Proceedings of the National Academy of Sciences (PNAS).
M. tuberculosis is dangerous because it hides and persists in the immune cells of our bodies. “It can only persist there because of the activity of key molecules,” says Matthias Wilmanns, Head of EMBL Hamburg. “We are investigating the functions of tuberculosis proteins and determining their atomic structures, in hopes of finding weak points and new inhibitors.”
A protein called LipB is essential for the organism because it activates cellular machines that drive the bacterium’s metabolism. Stefan Kaufmann’s department at the MPIIB has specialised in the biology of M. tuberculosis infection and its ability to survive in immune cells. They discovered that LipB is highly active in acutely infected cells, particularly in patients infected by multidrugresistant forms of M. tuberculosis.
“In these cells we see a 70-fold increase in the production of LipB when compared to other cells,” says Stefan Kaufmann,Director at the MPIIB.”This strongly indicates an involvement in pathogenesis and makes it a particularly interesting target where traditional drugs have lost their efficacy.”
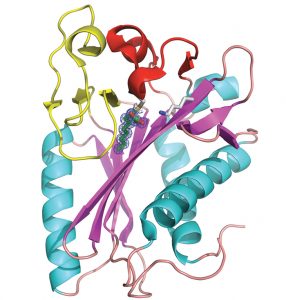
A structural picture of the protein – a kind of technical diagram of its building plan – has yielded important clues about its activity. Qingjun Ma from Wilmann’s group purified LipB and obtained crystals of the molecule. Using the high-energy synchrotron radiation beamlines at EMBL Hamburg, on the campus of the German Electron Synchrotron Radiation Facility (DESY), he created an atom-by-atom map of the protein’s structure. A highresolution picture of the active site of LipB bound to a lipid inhibitor helped to determine the function of the enzyme. In collaboration with EMBL’s Proteomics Core Facility in Heidelberg and researchers from the University of Illinois (USA) the Hamburg group discovered how LipB attaches specific fatty acids onto other proteins.
“LipB is a very promising drug target,” Wilmanns says, “because it belongs to a vital pathway. Unlike other organisms M. tuberculosis has no backup mechanism that could take over LipB’s role. This means that an inhibitor blocking its active site would shut down key processes the bacterium needs to survive and replicate. This would be a very effective strategy for a drug.”
The scientists will now search for compounds that can do so. At the same time, they are continuing to look for other proteins as drug targets. Wilmanns and his colleagues from various other institutes are now focusing on structures of molecules that help M. tuberculosis to persist in its dormant state and could become drug targets.
Over the past three years, EMBL has coordinated an “M. tuberculosis structural proteomics” consortium, supported by the German Ministry of Education and Research (BMBF), and produced high resolution images of more than 30 proteins. “Structure-based drug discovery has been a big success in the battle against many other diseases. We are now applying these tools to tuberculosis, one of the most devastating infectious diseases of mankind,” Wilmanns concludes.




