
Finding the way with X-Ray
New technique uses X-rays to find landmarks when combining fluorescence and electron microscopy
Imagine that your dog buried your wedding ring in the backyard, but you do not know where. Scientists face a similar challenge when they want to analyse individual cells located deep within tissues. Fluorescence microscopy allows researchers to monitor large samples, but it does not show the very fine details of cells. For this, scientists must turn to other methods such as electron microscopy. In many cases, the ideal solution would be to combine the two approaches, but the difference in scale can make this very complicated; like identifying an excavated patch in an aerial photograph of the backyard, then trying to find the same spot by examining close-ups of the ground inch by inch. A team of researchers led by Yannick Schwab, of EMBL, and Jacky Goetz, of Inserm in Strasbourg, developed a new approach that helps navigate between these different scales.
“Finding at the electron microscope something that you spotted under the fluorescence microscope is a tedious and time-consuming task,” says Schwab. “Not only is there a thousand-fold difference in the field of view of these two microscopes, but the sample distortions induced by the preparation for electron microscopy make it very difficult to target the correct spot with precision.”
Matthia Karreman, an interdisciplinary postdoc in the Schwab, Lénárt, and Hiiragi groups, was determined to make this task easier. Working with collaborators in Strasbourg, Heidelberg, and Munich, Karreman developed an approach that uses a third method, called microscopic X-ray computed tomography, to identify landmarks such as blood vessels in a sample.
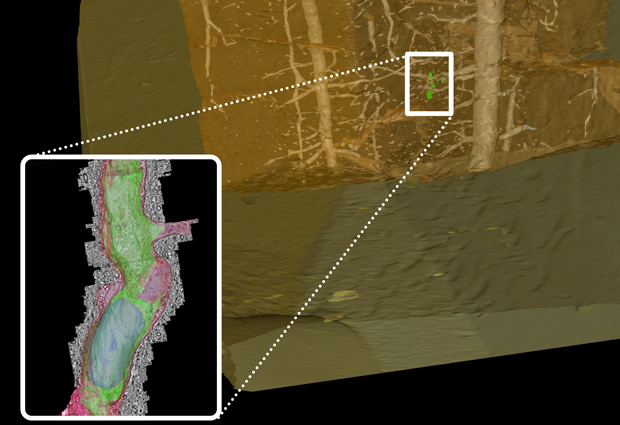
Like the X-rays doctors use to see inside our body without surgery, this technique employs X-rays to look inside the resin block in which a sample is embedded for electron microscopy, without cutting into it. Views from many different angles are combined using powerful computational tools to build a 3D image of the tissue inside the resin block. By comparing this 3D image with the one acquired at the fluorescence microscope, scientists can use the network of blood vessels visible in both images as a roadmap to find the cell of interest inside the resin block. They can then trim the resin block to expose the cell, and analyse only that fraction of the sample at the electron microscope.
“This approach makes it easier and faster to move from fluorescence to electron microscopy, because we know exactly which part of the sample to look at, instead of having to search through the whole sample for a marker,” says Karreman. Karreman, Schwab and collaborators have used their new technique, called multimodal correlative microscopy, to look at tumour metastases in the brain and skin of mice. Together with Péter Lénárt’s lab at EMBL, they also found tiny structures called centrioles inside the vastness of a starfish egg cell, which is a thousand times bigger than the average human cell.
Instead of analysing the whole cell in search of these minuscule structures – about three months’ worth of work – the new technique meant the researchers could analyse just the portion they were interested in, in around three weeks. “It’s my favourite type of technological development, because it is not a super new electron microscope, but it is a ‘trick’ that just helps speed up the experiment. As a biologist, that’s what I’m after,” says Lénárt. “We are very excited that other scientists are starting to use our technique to address their favourite questions in cell and developmental biology,” Karreman concludes, with a proud grin.


