
New mechanism for cancer gene activation
EMBL study using publically available data sets finds that rearranging how DNA packs the nucleus can activate cancer genes
A team of EMBL scientists has identified a new molecular mechanism for cancer gene activation using publicly available data sets. The team, led by Jan Korbel, found that rearrangements in the way DNA is folded within the nucleus can activate genes that would otherwise be silent, causing cancer.
DNA is formed in strands, long chains of molecular units that store information needed for life. Inside every cell nucleus we have around 2m of DNA, which is tightly twisted and looped around proteins to squeeze into such a small space. This means that particular sections of DNA can be physically close to one another even if they aren’t next to each other in their linear sequence. These are called topologically associating domains, or TADs.
TADs are up to around a megabase long, that’s about a million DNA bases, and if a gene and an enhancer are in the same TAD they can interact and the expression of a gene (the production of what the gene codes for) can be increased.
But there are ways in which the genome can be reorganised, altering its 3D structure. Korbel and his team used publicly available databases of cancer genomes and expression profiles to look at genes that are activated in cancers. Then they looked at the sequences and found something previously overlooked: in several human cancers, including colorectal and lung cancer, those genes involved in causing the disease are switched on by 3D structural changes to the DNA.
Characterisation through collaboration
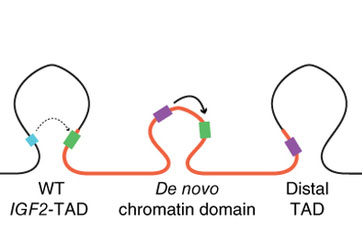
Taken from Weischenfeldt J et al (2016)
Although the team started with data sets and computational models, once they identified genes that they thought were activated in this way they contacted colleagues across Europe who had samples from colorectal and lung cancer patients. They then used molecular approaches to verify their model in these patient samples and characterise the mechanism of cancer gene activation.
Until now it has been a mystery how this cancer gene becomes activated
In one example, a large section of DNA encompassing a gene called IGF2 is duplicated. As the genome rearranges itself in space a new 3D structure emerges in the DNA that allows the IGF2 gene and a DNA regulatory region, termed enhancer, to come very close together. And so IGF2, which is known to be a cancer gene, is activated.
“You have to understand the molecular mechanism to understand why the cancer gene is so active,” explains Korbel. “People knew for a long time that IGF2 is an important gene in cancer, but until now it has been a mystery how this cancer gene becomes activated. It boils down to one mechanism, one gene.”
But without the data and computational analysis, adds lead author Joachim Weischenfeldt, now at the University of Copenhagen, that mystery would remain unsolved. “You need a lot of data to be able to identify such mechanisms.”


