
Mikhail Savitski
Team Leader, Senior Scientist and Head of Proteomics Core Facility
ORCID: 0000-0003-2011-9247
EditEmpowering research through mass spectrometry-based proteomics

Team Leader, Senior Scientist and Head of Proteomics Core Facility
ORCID: 0000-0003-2011-9247
EditThe Proteomics Core Facility provides a full proteomic infrastructure for the identification, quantification and characterization of proteins. This includes various platforms for protein and peptide separation, and state-of-the-art mass spectrometry for LC-MS/MS experiments.
Proteomics Core Facility
EMBL Heidelberg
Meyerhofstraße 1
69117 Heidelberg
Germany
Tel: +49 6221 387-8388
Email: pcf@embl.de
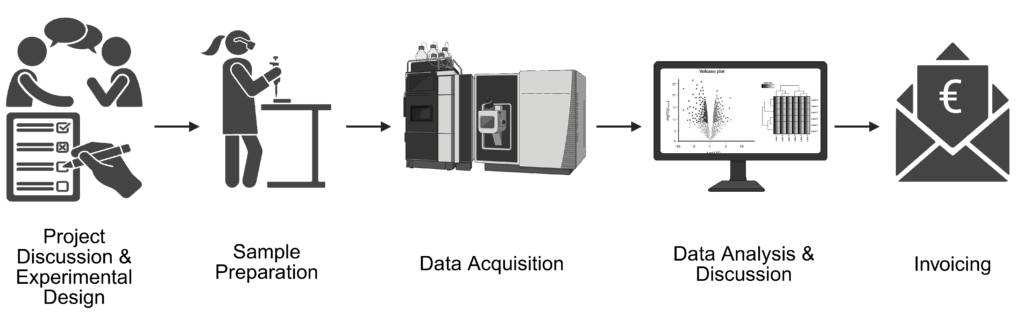
The Proteomics Core Facility offers a range of techniques for the identification, characterization and quantification of proteins and proteomes. These include the following techniques and approaches, often used in combination:
Please contact us for additional applications that may fit your needs.