Olivier Duss
Group Leader
olivier.duss [at] embl.de
ORCID: 0000-0003-0929-3471
EditAssembly mechanisms and function of protein-RNA complexes at the single-molecule level

Group Leader
olivier.duss [at] embl.de
ORCID: 0000-0003-0929-3471
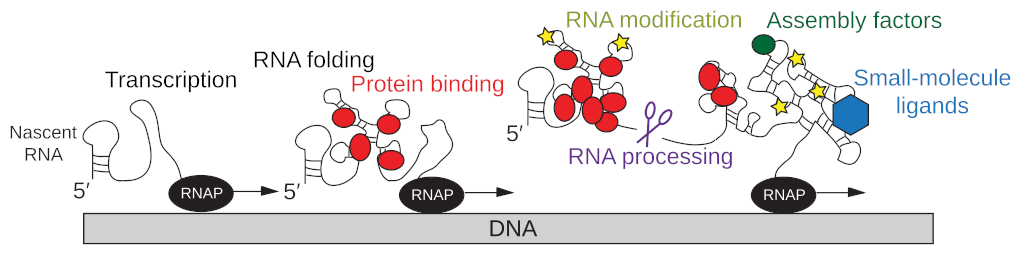
EditGene expression is coordinated by multiple interconnected processes including transcription by the RNA polymerase, simultaneous co-transcriptional RNA folding, RNA modification and RNA processing, the dynamic interaction of the nascent RNA with proteins and other ligands, and the translation of the mRNA into proteins by the ribosome (Fig. 1; Qureshi & Duss, FEBS letters, 2023). For some of these processes structural information is available, but we critically lack an understanding of dynamics and therefore the mechanisms of how these processes are functionally coupled to each other.
The challenge: While studying these mechanisms in vivo provides physiological relevance and an environment where coupling between the processes can occur, the cellular complexity makes it difficult to dissect the (dynamic) role of individual components in these interconnections. In contrast, in vitro reconstitution experiments provide a controlled environment allowing the manipulation of each component in the system but they often do not recapitulate the in vivo situation entirely because of their limited complexity.
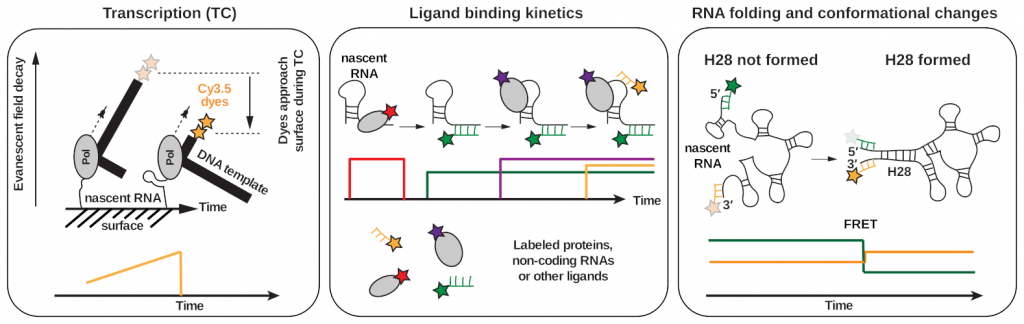
Our approach: Our lab bridges the gap between controllable, quantitative and high-resolution experiments from in vitro reconstitutions with in vivo physiological relevance. We reconstitute complex and active transcription, translation, mRNA processing & modification, and DNA repair machineries and investigate whether their activities and behaviors differ when studied in isolation versus when all components are present and the processes are simultaneously active. In order to directly visualize how various processes central to gene expression cooperate with each other, we take advantage of the recent explosion of high-resolution cryo-electron microscopy and cryo-electron tomography structures, representing numerous static states of different complexes involved in transcription, RNA processing & modification, translation, and DNA repair and use cutting-edge multi-color single-molecule fluorescence methods (Fig. 2) to place these various structural snapshots along a reaction trajectory. Together with integrative structural biology, biochemistry and in vivo single-molecule tracking, this combination of methods (Gor & Duss, 2023) allows us to dissect how various macromolecular machines cooperate and work in context.