Submit a TREC sub-project
Join in!
If you would like to participate in the TREC project with your samples, data, or analysis, please write to us: TREC@embl.org. Download template (Word)
Traversing European Coastlines
An expedition to study coastal ecosystems and their response to the environment, from molecules to communities
The TREC expedition aims to deepen our understanding of coastal ecosystems and how they respond to environmental change at the molecular level. By uncovering the mechanisms that drive ecosystem function and resilience, TREC is generating new knowledge – and potential solutions – for addressing the harmful environmental impacts of human activity on coastal ecosystems.
TREC projects are highly collaborative, bringing together EMBL researchers and those from partner institutions. See above the expanding TREC projects portfolio with sub-projects submitted by the growing network of collaborators.
Join in!
If you would like to participate in the TREC project with your samples, data, or analysis, please write to us: TREC@embl.org. Download template (Word)
Since the project list is long, we recommend using your browser’s search feature. Press Ctrl + F (or Cmd + F on Mac) and type in a keyword to quickly find projects of interest.
This project aims to assess how microbial biodiversity is impacted by sea, freshwater and estuaries, pollution, and other environmental factors. In the Bork lab at EMBL, for example, one of the focuses is on the gene flux between land and water. More specifically, soil is a major reservoir of antimicrobial resistance (AMR) and this project will study local AMR profiles and mechanisms of AMR spreading from soils to waters and humans.
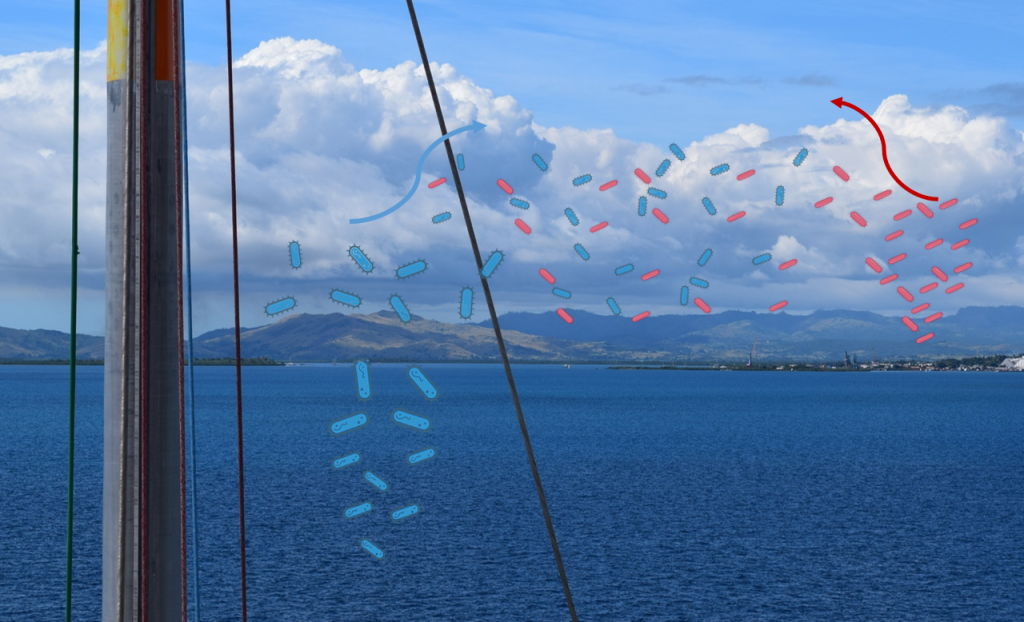

Bioaerosols remain one of the least studied microbiomes on Earth. Airborne microbes impact the health of humans, plants and animals, and drive key climatological processes. The TREC project enables the Sunagawa lab at ETH Zurich to team up with the Ocean-atmosphere interactions group, led by Michel Flores at the Weizmann Institute of Science, and explore the diversity, function and dispersal of airborne microbial communities at the land-ocean interface. (BIOcean5D)
The Brunet lab at the Institut Pasteur (Paris) studies choanoflagellates, the closest living relatives of animals. Many choanoflagellates can switch between unicellular and multicellular forms and thus offer a proxy to the origin of animal development. However, only a fraction of existing species are known. We will try to systematically describe and isolate new culturable choanos from plankton to understand the natural environment for the emergence of multicellularity.
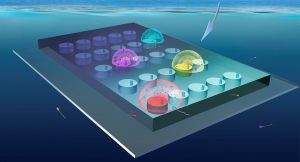

Microbial behavior will be quantified during TREC through the deployment of a novel sea-going microfluidic platform, the In Situ Chemotaxis Assay (ISCA). This international collaboration (Stocker: ETH Zurich, Raina + Seymour: University of Technology Sydney, Vincent: EMBL) will leverage the contextual data gathered during the expedition to identify environmental drivers influencing microbial behaviors.
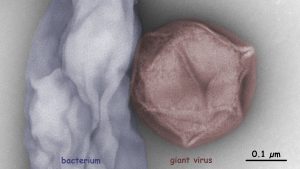
Giant viruses influence the mortality, population structure and gene flux of various protists. The Fischer lab at the MPI for Medical Research studies the diversity and biogeography of giant viruses by electron microscopy, metagenomics, and cultivation on heterotrophic flagellates. We aim to compare coastal virus communities from water and soil and to isolate new giant viruses for a better understanding of their ecology.
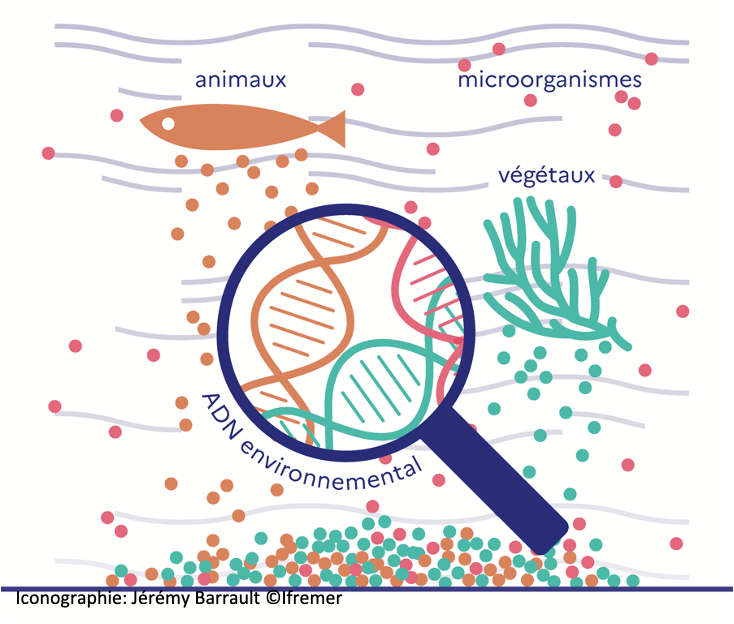
As part of the TREC expedition, Dr Arnaud-Haond (Ifremer) coordinates a team of researchers (AZTI, Ifremer, CNRS) to characterize marine communities across the entire Tree of Life (i.e. prokaryotes, protists, metazoans…), using metabarcoding protocols based on environmental DNA. Sampling performed along shallow-coastal gradients, under diverse levels of human pressure, will allow mapping the biogeography depending on life history traits and assess the impact of human activities.
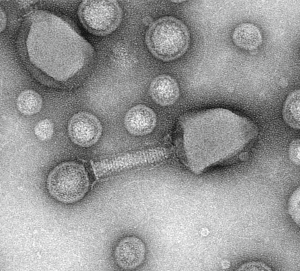
Viruses and Vesicles (VV) significantly shape plankton communities and marine biogeochemical cycles. Host-virus-vesicles interactions are poorly characterized although mediating key processes such as transfer of nucleic acids, antibiotic resistance, and cross-kingdom cell-cell communications. The Plankton Vesiclomics project (Chaffron, Baudoux et al., CNRS) will characterize the diversity and ecology of host-virus-vesicles interactions in coastal ecosystems by combining imaging, viral particles/vesicles quantification, sequencing, and computational analyses.
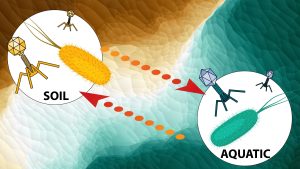
The Sullivan Lab at the Ohio State University will examine, through virus-enriched metagenomics and linked microbiome datasets, the diversity and impact of viruses that infect soil and aquatic microbes. Virus abundance best predicts carbon fluxes in global oceans and impacts the evolutionary trajectories of their hosts. However, we lack data on such dynamics in coastal systems, which could be key in devising efficient climatic and ecological models for these areas.

Ifremer will analyse microbiome and environmental data from the shallow waters of the river outlet stations during the TREC expedition to identify biodiversity patterns of estuarine microbiomes and the factors structuring this diversity. Spatial data will be integrated to the ROME eDNA national observatory network to validate at temporal scale the potential differences found between estuaries of different basins – English Channel, Med. Sea, Atlantic Sea. (BIOcean5D).
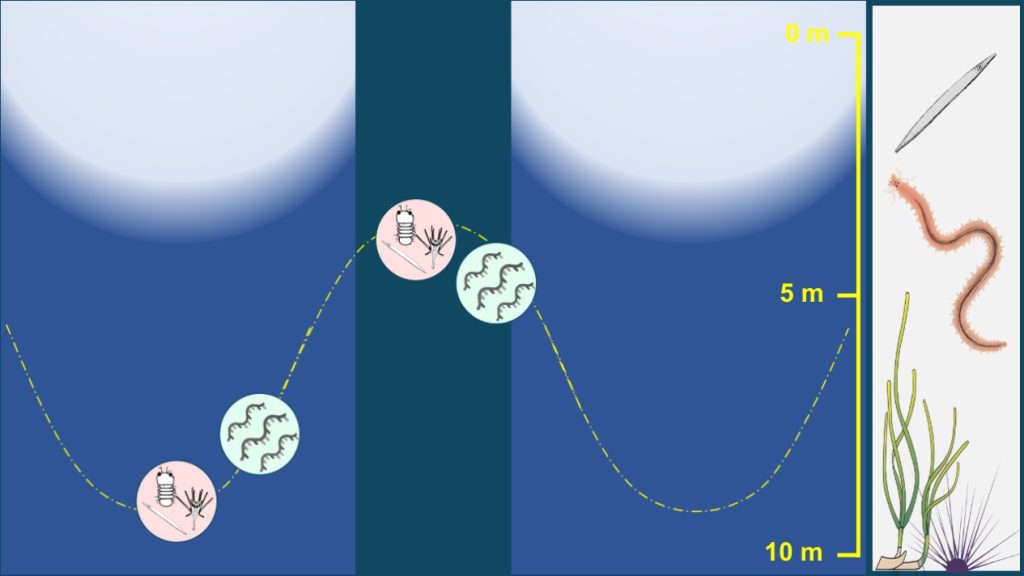
This project, lead by SZN (ITA) in collaboration with EMBL (GER) and other groups (in AUT, FRA and GER), aims at providing a multiscale analysis of plankton and benthos dynamics during the light-dark cycle in a coastal area of the Gulf of Naples. Meta-barcoding, meta-transcriptomes, morphological analysis and physiological measurements will be combined to describe diel rhythms at sea at the ecosystem level.

The Complex Team (LOV, Sorbonne University, Villefranche sur Mer- France) will analyze using the Flowcam the samples collected by 20µm nets. Results will allow us to quantify and qualify the microplankton across European seas and provide insight into the plankton ecology and trophic ecology in the various environments sampled.

The Complex Team (LOV, Sorbonne University, Villefranche sur Mer- France) will analyze using the Zooscan the samples collected by 200µm nets (WP-2 nets). Results will allow us to quantify and qualify the mesoplankton across European seas and provide insight into the plankton ecology and trophic ecology in the various environments sampled.
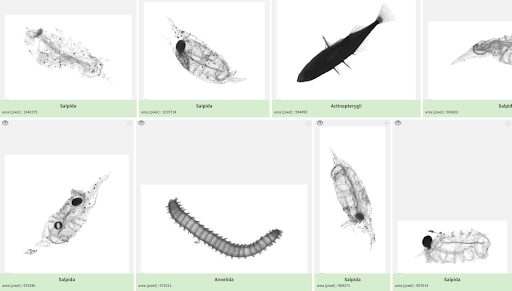
The Complex Team (LOV, Sorbonne University, Villefranche sur Mer- France) will analyze using the Zooscan the samples collected by 680µm nets (Regent). Results will allow us to quantify and qualify the macroplankton across European seas and provide insight into the plankton ecology and trophic ecology in the various environments sampled.
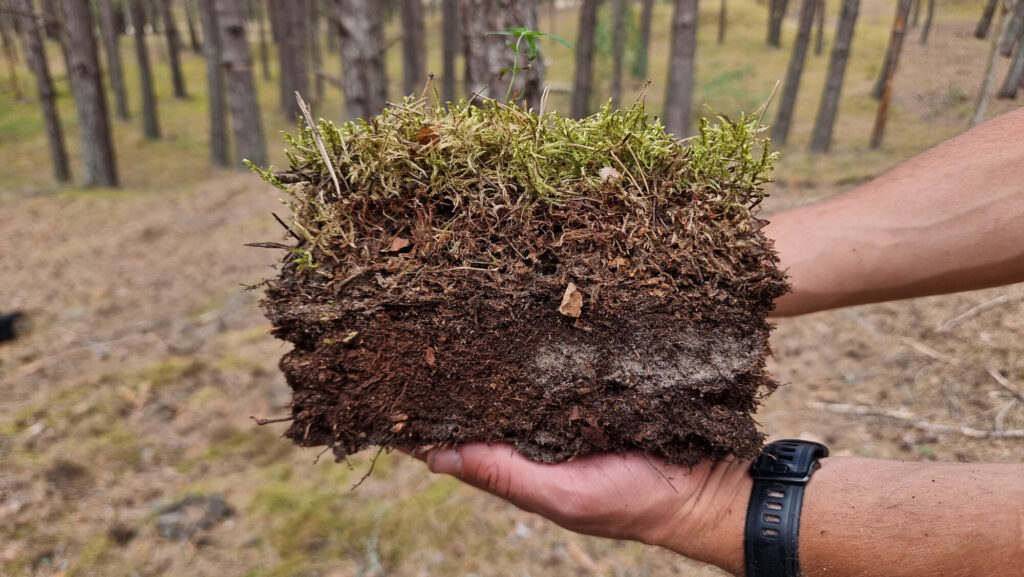

Archaea represent the third domain of life; however, it is considerably understudied. Originally believed to only exist in extreme environments, they have since been shown to be ubiquitous in the environment. By using metagenomic techniques the Bork group at EMBL can explore the extent of archaeal diversity and their roles in the environment.

The Flores lab at the Weizmann Institute of Science will explore the airborne plastisphere—microbes hitchhiking on microplastics in the air—along 14,000 km of European coastline. By characterizing both the plastics and their microbial communities, we aim to understand their diversity, transport potential, and implications for ecosystem and human health.
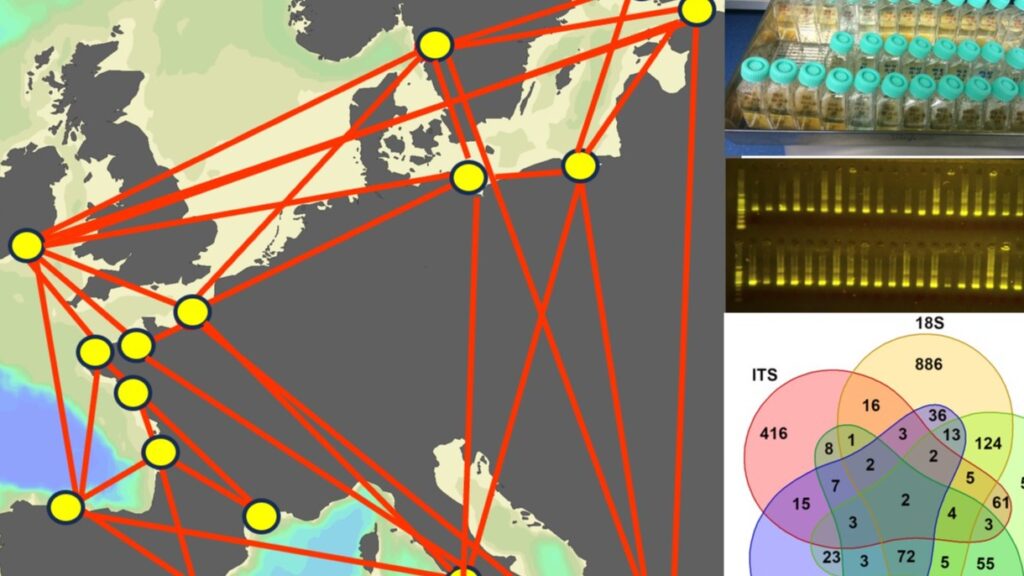
Genetic variability within species forms a basis for responding to environmental selection and provides information on the connectivity among geographically separated populations. The Phytoplankton group at TalTech, led by Prof. Sildever, will investigate the population genomic structure of cosmopolitan harmful algal bloom species using SNPs to understand their capability to spread in European coastal waters.
Mobilome is an overarching term describing mobile genetic elements of different nature (phages, plasmids, integrons, transposons) residing in the microbiome. The Bork group at EMBL developed a computational pipeline to discover microbial genes that have potential to be mobilized and horizontally transferred in microbial communities. In a changing world marked by rising temperatures, pollution, and human impact on land and sea, the composition and function of mobilised genes play a critical role in enabling adaptive responses. TREC expedition gathered crucial data that enables unprecedented resolution in identification of environmental drivers of gene mobility that can help to identify key factors contributing or restricting spread of microbial adaptation traits e.g. resistance to antibiotics or pollutants degradation.
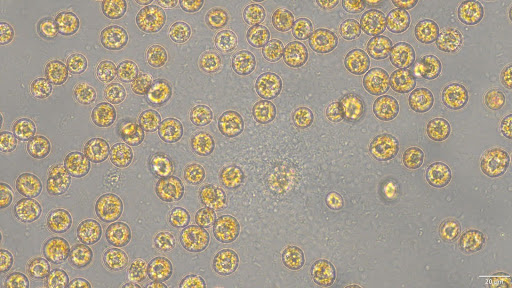
Measuring the taxon-specific lytic mortality is a crucial challenge to understanding the biogeochemical cycles. The Ogata lab at Kyoto University will investigate the extracellular rRNA of microeukaryotes to address their cell lysis in the environments. We aim to reveal the organisms that are actively lysed and their relationship with environmental conditions.

VIROBS investigates the temporal dynamics of marine viruses in the Western English Channel. Combining molecular and biogeochemical approaches, the lab of Dr Baudoux (CNRS) aims to monitor viral abundance, diversity, and activity to better understand virus-host interactions and their impact on carbon and nutrient cycling under environmental pressures.
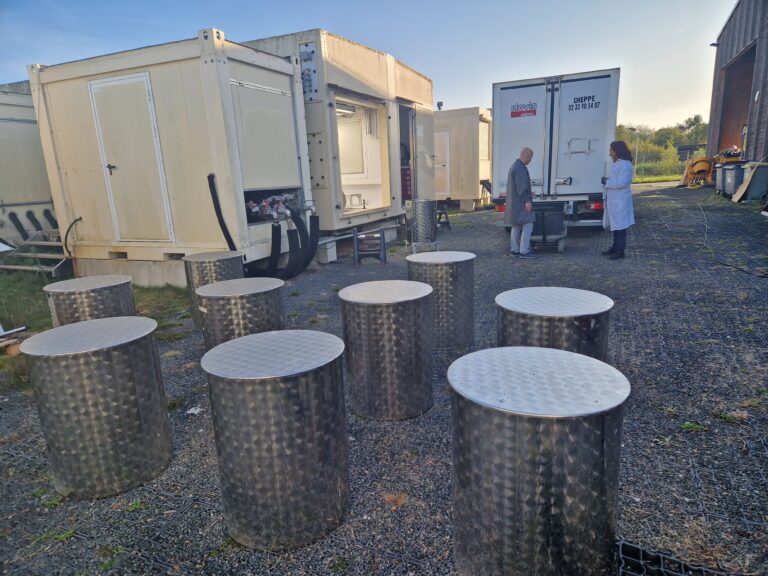

The project led by the Bork and Zimmermann groups, unites the molecular biology expertise of EMBL with the high-tech mesocosm facilities and knowledge of Ecotron Ile de France to tackle questions on the microbial changes and adaptation in global warming and human-induced pollution. The derived results will generate new knowledge about how coastal ecosystems respond to anthropogenic pressures such as climate change and xenobiotic pollution.
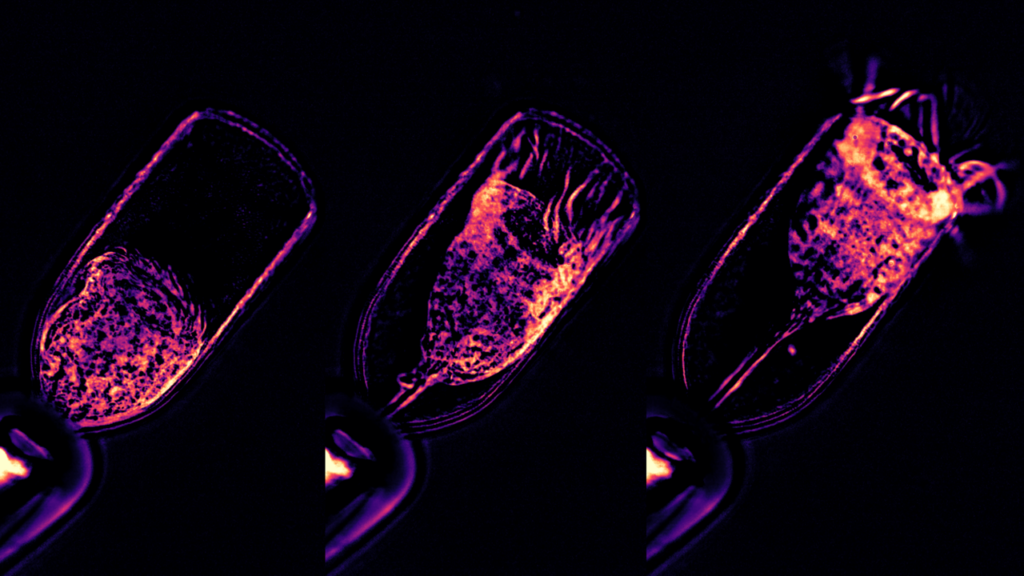
Protists have surprisingly cognitive behaviours, such as mating, feeding, and prey capture. The Wan lab at the Living Systems Institute (University of Exeter) will chart the motility of diverse marine protists and their responses to sensory stimuli. The aim is to correlate motor outputs and excitable behaviours with cellular structure, to discover how functional complexity may be embodied within the physical architecture of these organisms.

The AtlaSymbio project aims to improve our understanding of aquatic photosymbioses by unveiling the three-dimensional ultrastructure of host-microalgae interactions using 3D subcellular imaging. We will generate an open-source atlas capturing the architecture and metabolic implications of different photosymbioses collected during the TREC expedition or with experts. The ambition is to foster collaborative exploration and advance the mechanistic understanding of aquatic photosymbioses.
AtlaSymbio is funded by the Gordon and Betty Moore Foundation. The project involves an interdisciplinary consortium of experts in microalgae, symbiosis and electron microscopy, from Grenoble (Decelle’s group, Cell and Plant Physiology Lab-Photosymbiosis team), EMBL Heidelberg (Schwab’s group), and Roscoff (Ian Probert, Marine Station, Roscoff culture collection).
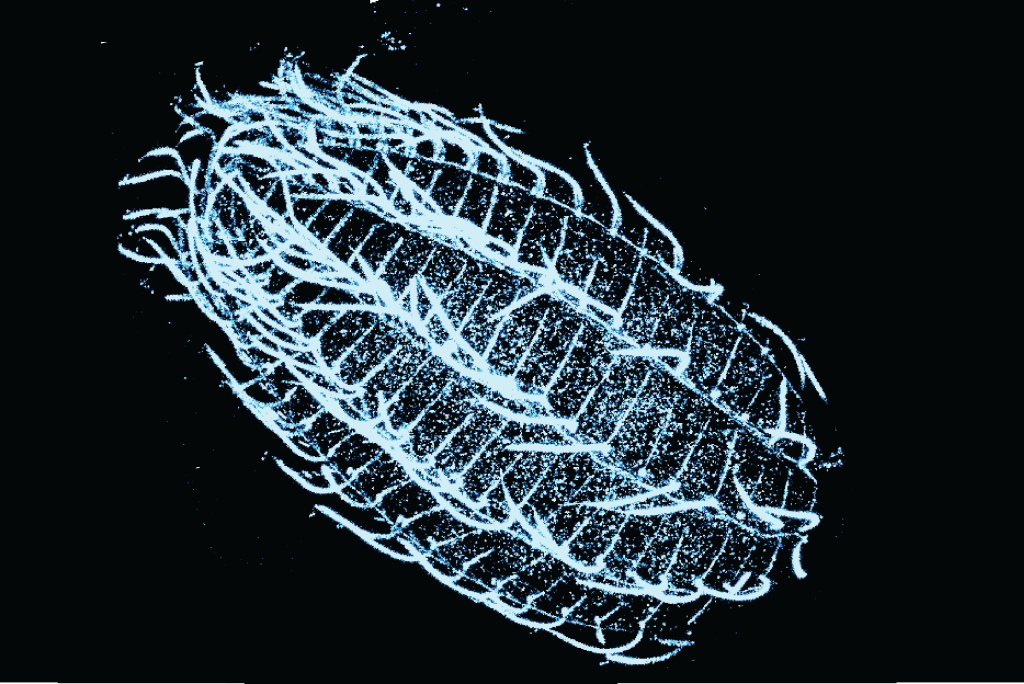

An international EMBL-EPFL-UNIGE team (Dey, Schwab, Saka, Dudin, Hamel, Guichard) are coupling expansion microscopy (Cryo-ExM and U-ExM) to FISH for species ID and sub-cellular imaging. We aim to create an atlas of native eukaryotic cellular biodiversity with unprecedented 3D spatial resolution, with a long-term view to assess the repercussions of climate change on free-living microplankton populations.
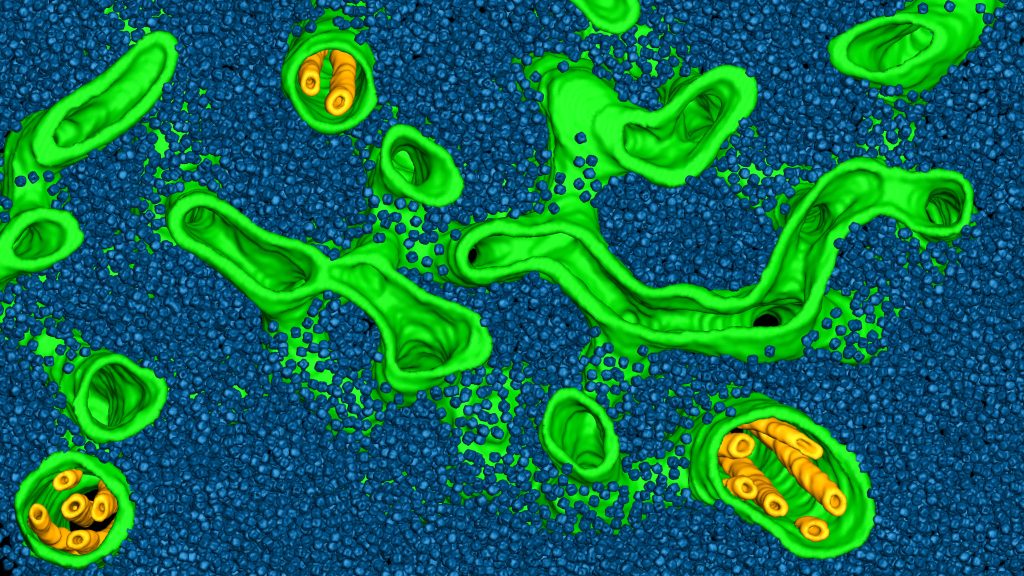
The Engel group at the University of Basel uses cryo-electron tomography (cryo-ET) to visualize structures inside native cells with molecular resolution. In the cryOcean project, we aim to chart the molecular organization of photosynthetic organelles (light-harvesting thylakoid membranes and carbon-fixing pyrenoids) in diverse marine algae species sampled directly from the ocean.
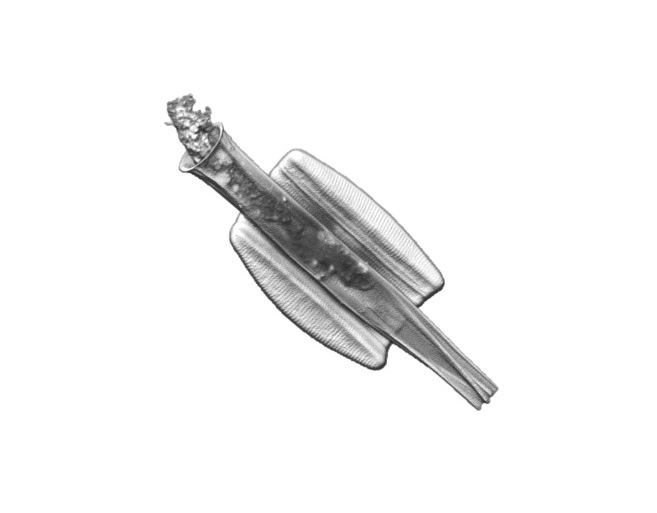

This project is dedicated to understanding the diversity, distribution and impact of intimate interactions involving diatoms, major primary producers in the ocean. By using a combination of single-cell live and subcellular imaging, high-throughput phenotyping approaches, and in situ chemotactic assays, the Vincent lab at EMBL asks how microbial interactions can affect diatom behaviour, development and survival in a changing ocean.

NAVARRO aims to capture dynamic cellular processes in diverse marine environments – bridging structural biology with environmental studies. With advanced cryo-EM sample preparation instruments now available directly at the marine sampling sites, The Schwab Lab at EMBLwe have begun investigating molecular machines in situ, within their natural context.
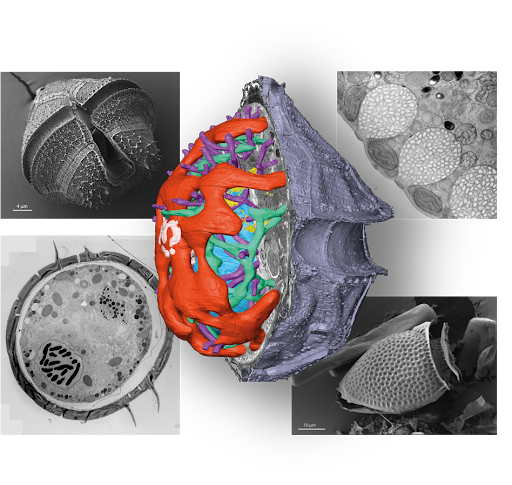
The Schwab Lab at EMBL employs high-resolution correlative electron microscopy to reveal the complex cellular architecture of marine dinoflagellates from natural environments. We are constructing a geo-centric ultrastructural atlas of dinoflagellates using high-resolution 2D EM, high-throughput X-ray tomography, and volume EM methods.
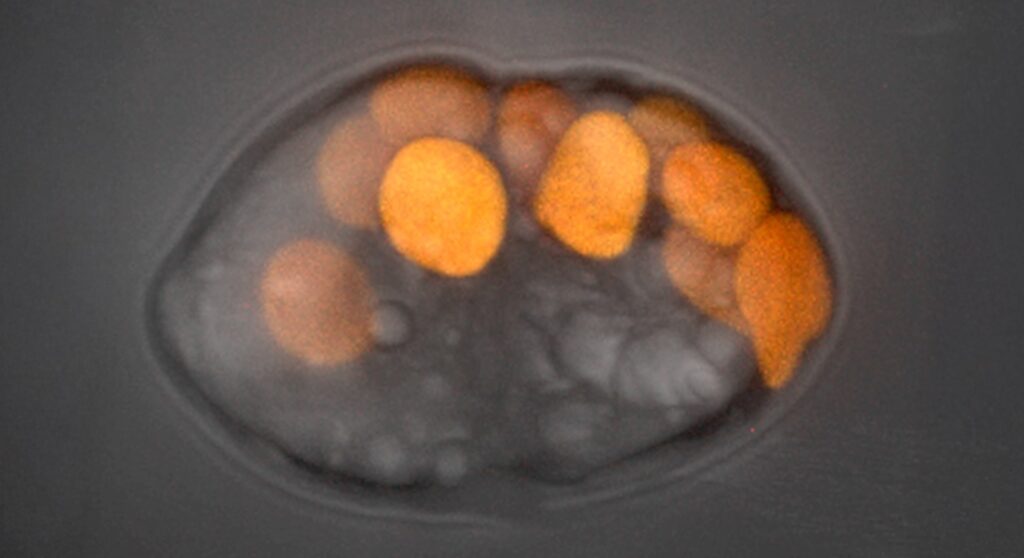
The Hehenberger Group (Biology Centre, CZ) investigates the processes during plastid endosymbiosis using microorganisms that steal and transiently retain plastids from their prey. We aim to analyze the physiological and genetic integration of the plastids in such lineages by combining cultivation-dependent and -independent approaches with the support of the AML. The project is part of the ERC CoG-funded project KLEPTOS.

Diatoms from genus Chaetoceros are known to form chains that can go up to 50 cells. Because of technical limitations, our knowledge of why those chains are formed is limited. Thanks to in situ single-cell sampling made during the TREC expedition and single-cell RNA sequencing, this project in the Vincent lab at EMBL, aims to explore the impact of environmental conditions on chain formation and length, and decipher the ecological importance of chains.
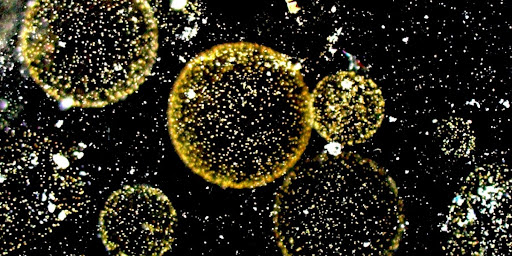
The main goal of the project, led by Decelle group at CNRS/UGA/CEA, is to better understand the life history of the marine microalga Phaeocystis, which is a keystone phytoplankton taxon in the ocean. In coastal waters, blooms of colonies can have negative impacts in the ecosystems with consequences on the fisheries and touristic activities. The ability to live in symbiosis intracellularly in a host cell is also an intriguing phenomenon in Phaeocystis. By unveiling the diversity of the microbiome across these different life stages and along the European coast, we hope to bring new information to better understand the ecological success of this microalga. The microbiome will be assessed by different sequencing strategies thanks to the samples collected during the TREC expedition. This project forms part of the BIOcean5D EU funded project.

Plankton drives marine ecosystem function, but phenotyping cells in their environment remains challenging. Manual sorting of morphologically diverse samples creates discovery bottlenecks. This TREC project, uniting EMBL, Roscoff Biological Station, and Ifremer Brest, advances image-enabled cell sorting across all planktonic size fractions. By integrating computer vision, photolabelling, and high-throughput single-cell sorting/sequencing, it aims to accelerate marine research capabilities.
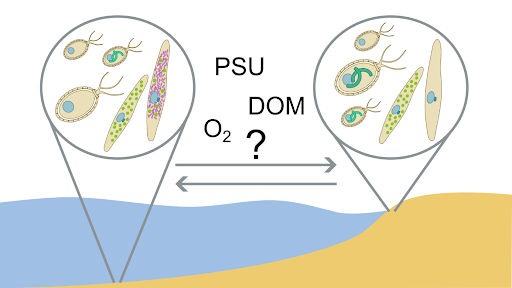
This project will analyse the TREC dataset to investigate species diversity and geographic distribution of two unique protist endosymbioses: Paulinella, the genus within which an independent photosynthetic organelle evolved, and Pseudoblepharisma, the genus within which the only known ‘green and purple’ endosymbiosis evolved. The team (Sørensen, Muñoz-Gómez, Nowack & Burki) will use this data to study the effect of environmental gradients on the endosymbiotic state of these protists. Ultimately, our goal is to explore how endosymbiosis-mediated metabolic innovation can enable habitat expansion.

Cosmopolitan organisms possess the ability to easily adapt to new conditions and habitats. The molecular changes orchestrating the adaptation and microevolution of particular cell types and tissues to changing conditions as in times of global warming are, however, poorly understood. The goal of the Arendt group at EMBL Heidelberg, is to identify cell types that are prone to environment-driven change for the cosmopolitan marine worm Platynereis dumerilii. We are currently sampling different P. dumerilii populations across European coastal sites to unveil transcriptomic differences within different cell types. We have sampled populations from Sweden, Spain, France, Italy and Greece which provide insights into abundance changes for certain cell types and site-specific gene expression patterns.
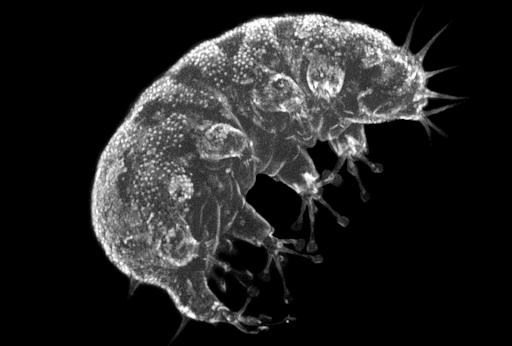
Meiofauna is a vital link in benthic ecosystems. Ifremer, in the context of Meiodyssea project (https://meiodyssea.ifremer.fr/) will characterize the diversity of meiofauna present in European coastal zones using imaging methods based on flow cytometry to understand the environmental and anthropogenic factors shaping these communities, on a scale unprecedented for these organisms.
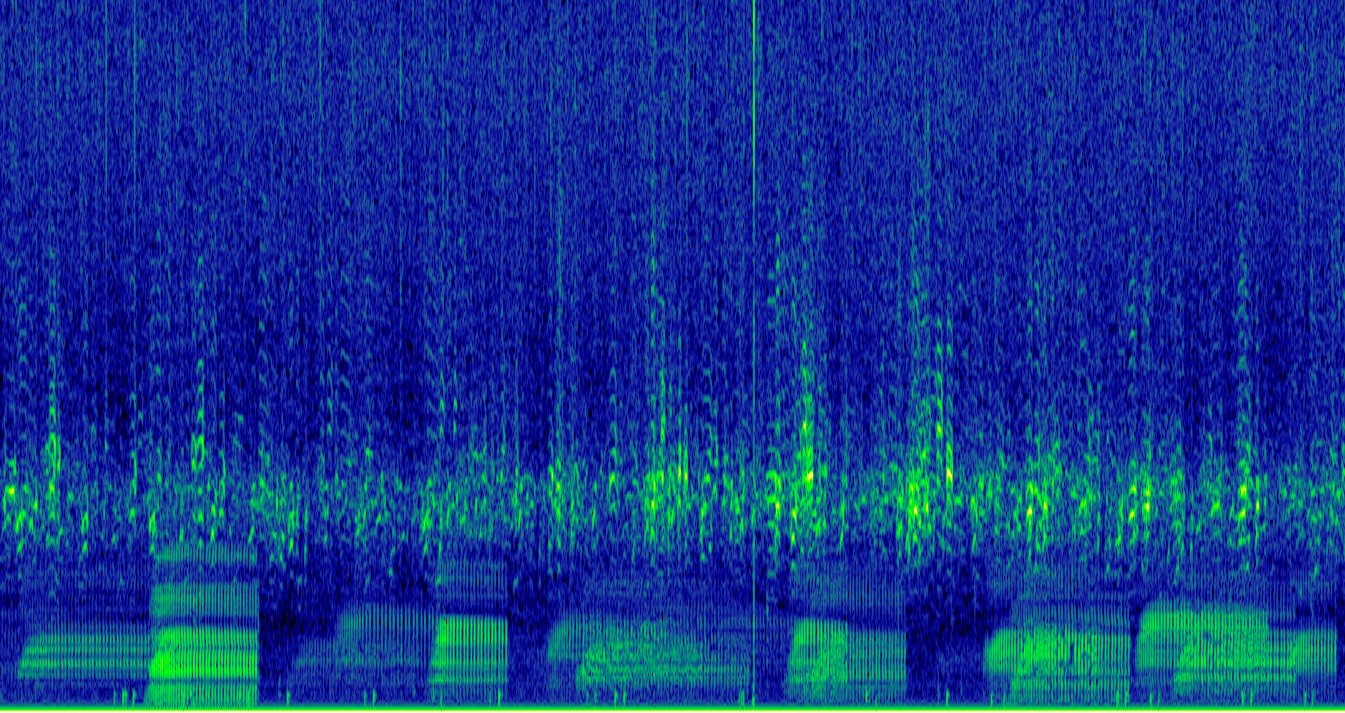
As part of the EU BIOcean5D project, L. Di Iorio and co-workers at UPVD will assess the biogeography of bioacoustic diversity and investigate environmental state-pressure relationships using soundscapes. Noise pollution will be quantified to study its impact on model systems. The collaboration with the CNRS & EMBL involved in other TREC projects will allow to appraise functional aspects of soundscapes.
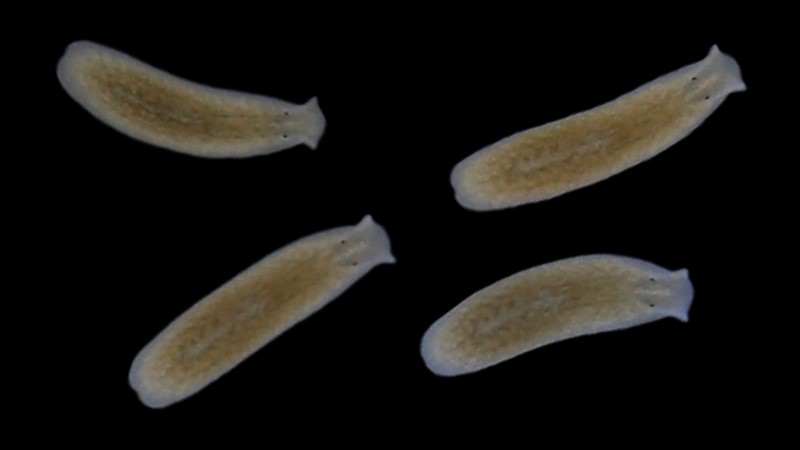
Why can some species live in a wide range of habitats, while others cannot? The Vu group (in collaboration with many folks at EMBL and beyond) aims to answer this question by first, carrying out a systematic field sampling of planarian flatworms along the coastline of Europe; and second, applying cutting-edge single cell multiomics and targeted perturbations to wild planarians brought back to the lab.
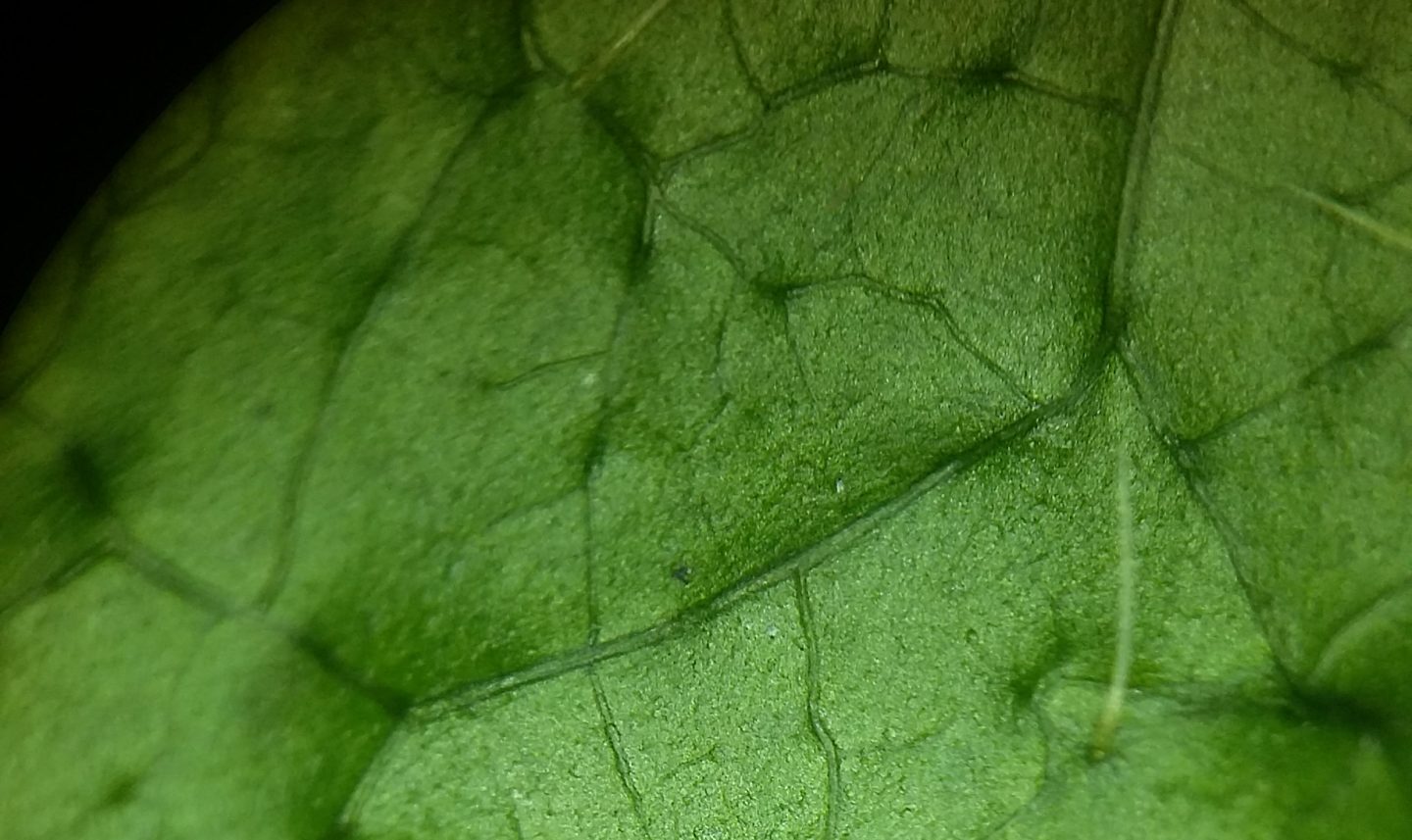

FunLeaf project is led by Dr. Niloufar Hagh-Doust at University of Tartu, Estonia. This citizen science project aims to describe the biodiversity of microorganisms associated with the plant leaves across the globe. Using DNA-based methods for identification, the aim is to determine the environmental features that shape the leaf microbiome and add one piece to the puzzle of global biodiversity.
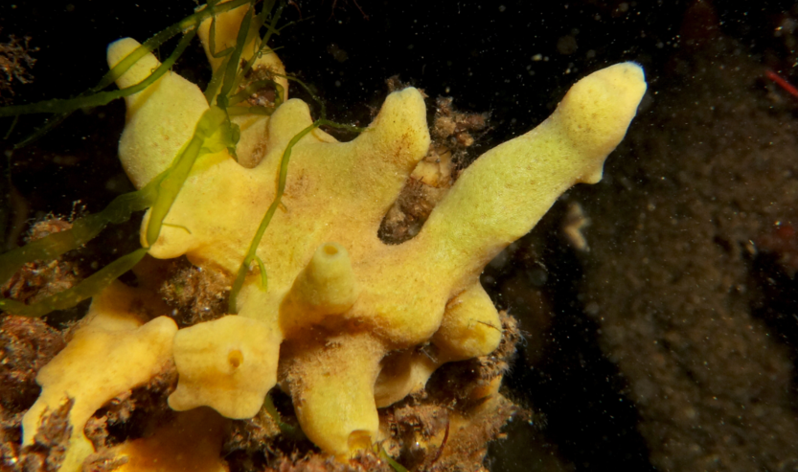

The Hentschel lab at GEOMAR will define the effects of environmental gradients on microbiome variation and individuality in the sponge H. panicea. We will explore taxonomic and functional gene variation of the dominant microbial symbiont clade and immune gene variation on the host side. Our overarching aim is to move marine sponge symbioses from an exploratory to experimental arena for host-microbe interactions.

Led by the Hambleton lab (Uni Vienna), this project will use transcriptomics and lipidomics to assess the biogeography and function of dinoflagellate algae in marine sediments and in symbiosis with marine invertebrates. We aim to reveal how key symbiotic organisms are affected along evolutionary and environmental gradients, allowing predictions of how these symbioses will respond to accelerating climate change. Part of the BIOcean5D EU-funded project.
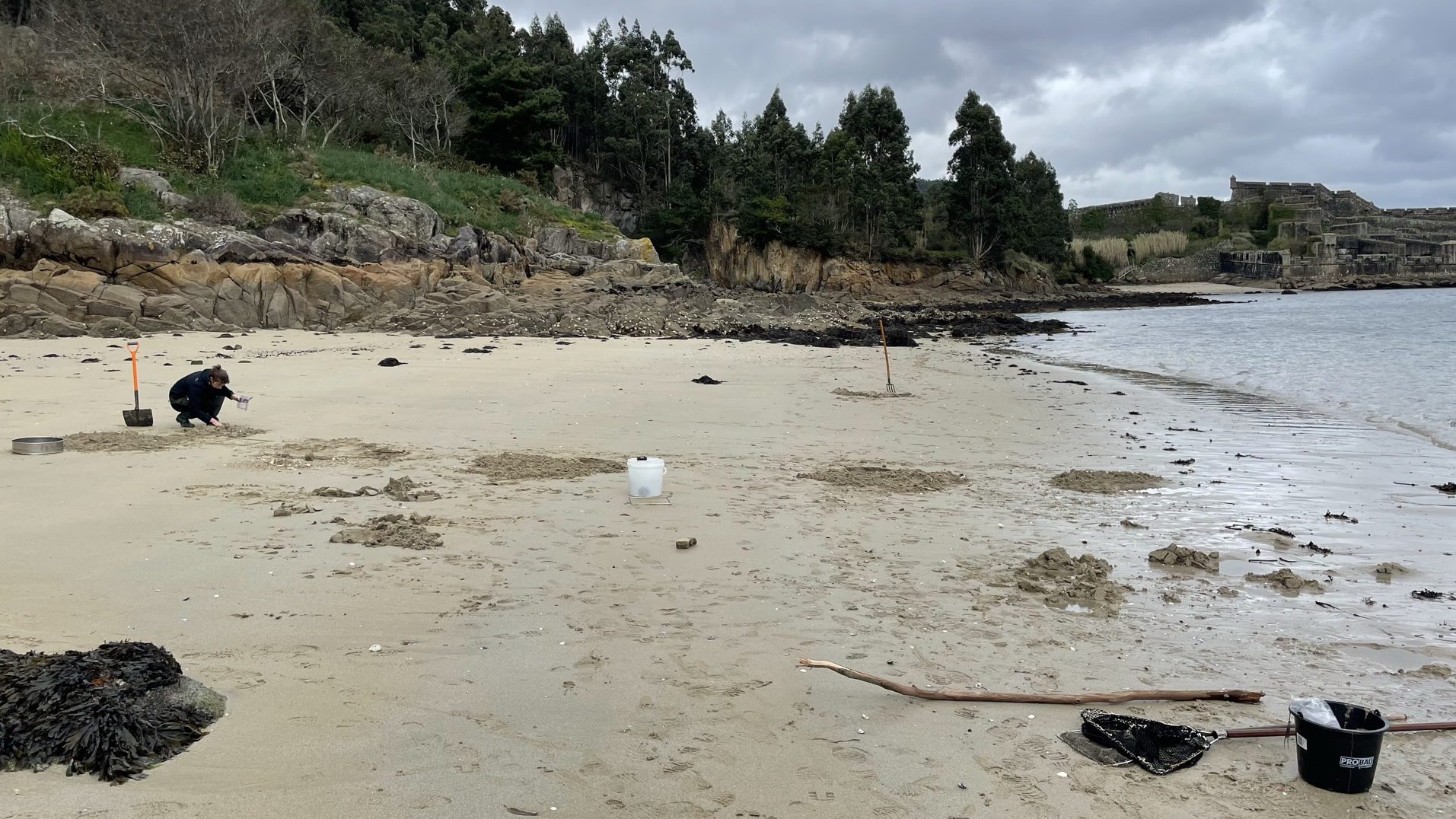

The Animal Evolution and Biodiversity department at the University of Göttingen investigates the annelid macrofauna across the European coast in TREC project. Animals are sampled across a defined transect covering the upper-, middle-, and lower-intertidal zones at sandy and rocky shores. A subset of focus taxa is analysed morpho-genetically to unravel taxonomic distribution patterns and to corollate it with the biotic and abiotic factors. (BIOcean5D)
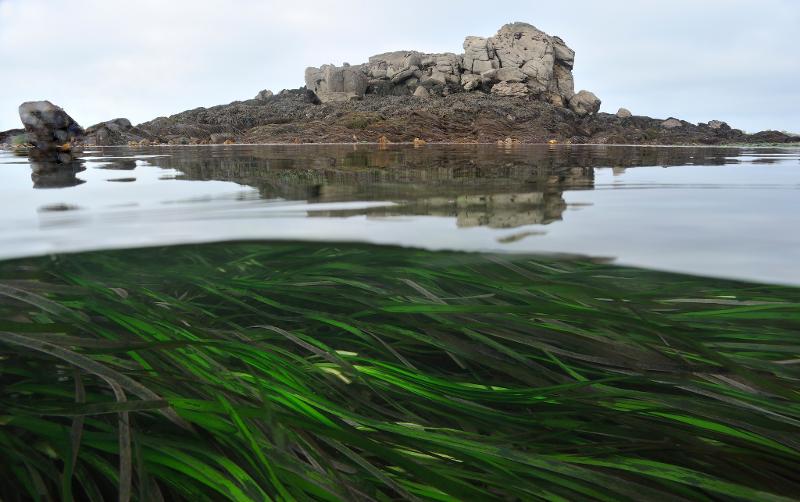

In the frame of the Biocean5D project, the teams of Jillian Petersen (University of Vienna) and Sophie Arnaud-Haond (Ifremer-MARBEC, Evolution and Genetics of Marine Organisms) will coordinate their expertise in microbiology and population genomics of clonal engineer species to characterize the ‘holobionts’ structuring seagrass meadows along European coasts, in order to map biotic interactions and reveal the environments conditioning this possibly fragile equilibrium.
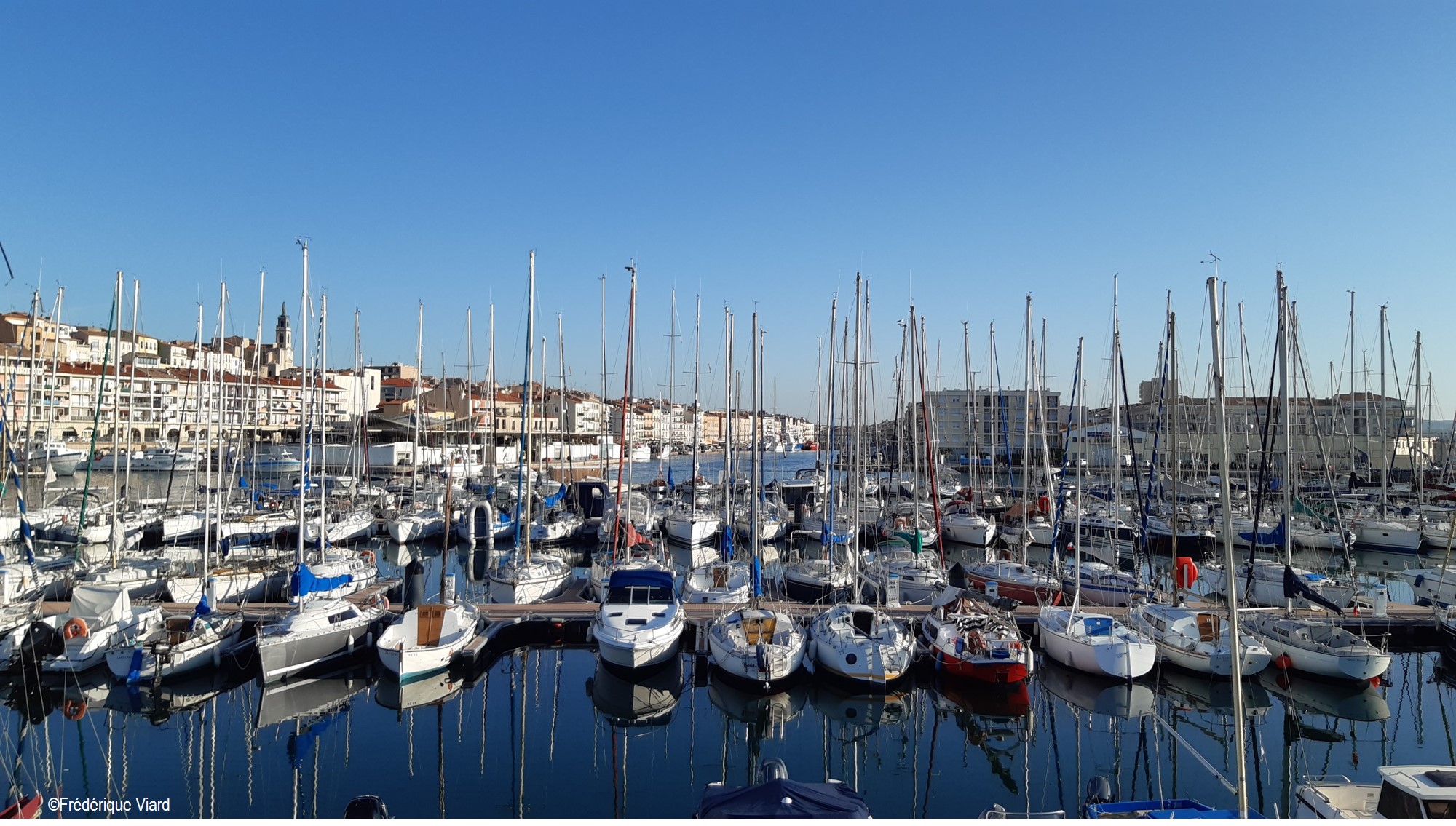

As part of the Horizon Europe BIOcean5D project, Dr Viard (CNRS) will lead a team of researchers (CNRS, Ifremer, CSIC) to identify, using techniques based on environmental DNA, non-native and invasive marine species present in ports and surrounding natural habitats. This project will allow assessing the importance and spread of non-native species from local to European scales, and will also inform on the role of harbors in these dynamics.
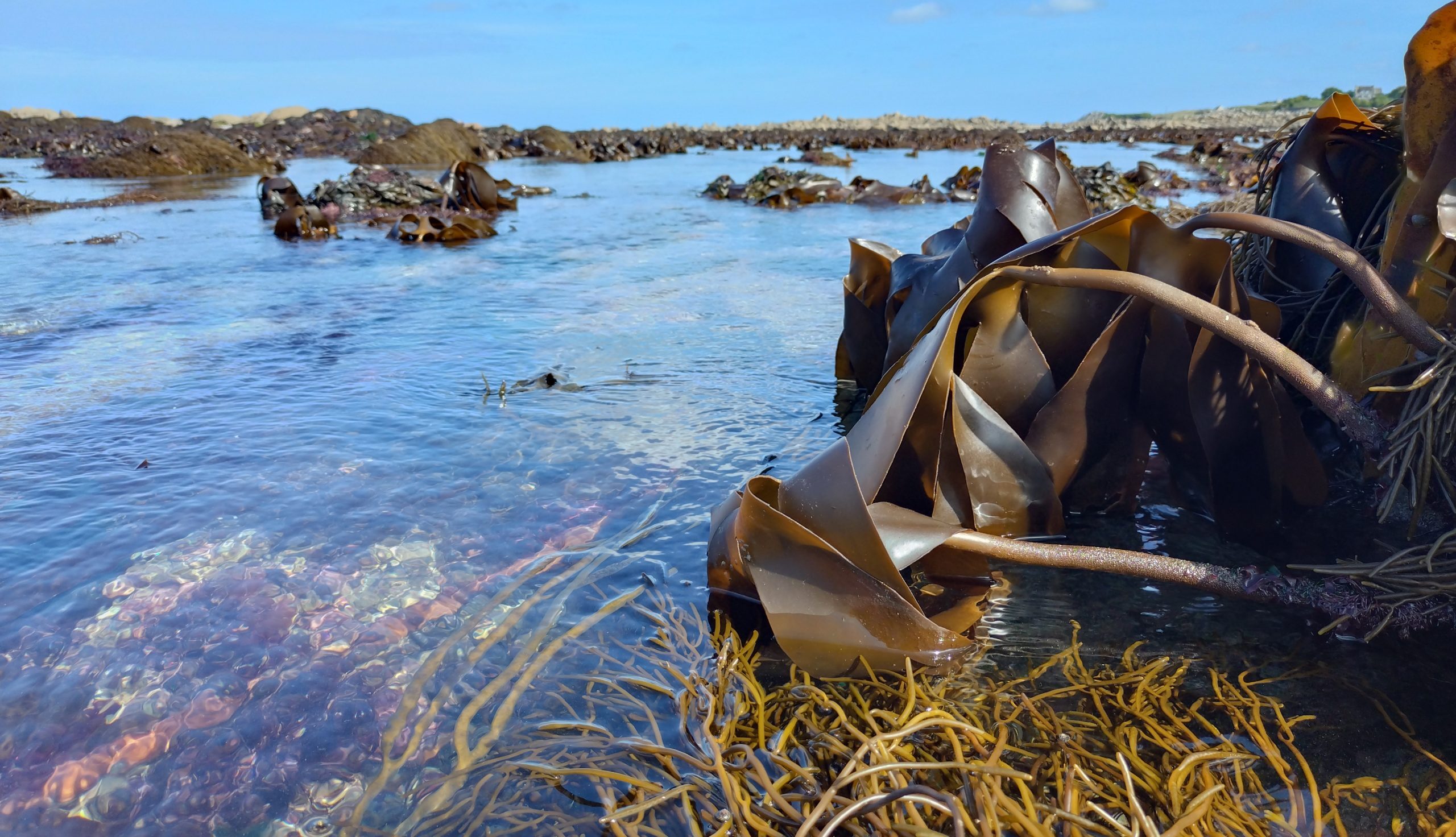

In this joint project between MPI Tübingen (GER) and SB Roscoff (FRA), we will study the biodiversity of kelp forests and their associated microorganisms in European coastal waters. We aim to link the genetic variation of two kelp species, Saccharina latissima and Laminaria hyperborea, with the diversity of their associated microbiota and viruses, focusing on intra- and inter-site variability with respect to the phylogeographic history of the populations. (BiOcean5D)
This project investigates the diversity and distribution of phytochemicals across European coastal ecosystems, focusing on how environmental gradients — such as edaphic factors and climate — influence the production of bioactive compounds in coastal plant species. By integrating high-throughput fine-scale phytochemical analyses with ecological data, the group of S. Rasman and E. Defossez at Université de Neuchâtel, Switzerland, aim to understand the adaptive strategies of coastal flora and their potential applications in biotechnology and medicine. The findings will contribute to conservation efforts and the sustainable utilization of coastal plant resources.

Sequencing genomes can reveal genes underlying adaptation to new environments. However, it is challenging to trace the function of a genetic change to its effect on organismal survival. Here, the Dorrity group will use a similar conceptual approach to instead profile cells in species adapted to different thermal environments, collaborating with the Arendt group to perturb cells and in genes associated with adaptation to natural thermal environments in lab populations of Platynereis.
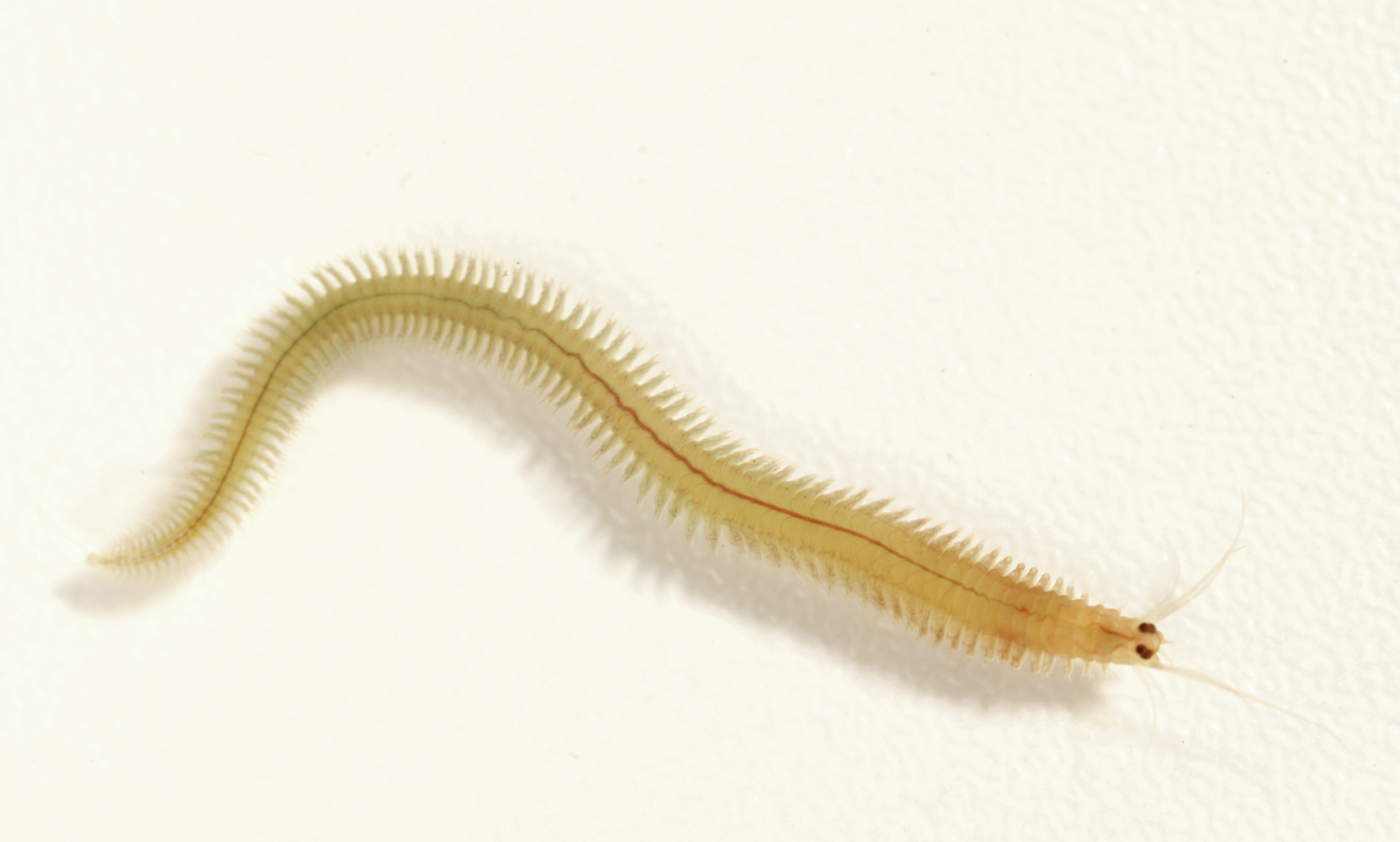

This project, led by the Arendt lab at EMBL and collaborators, will explore the cellular composition of the brains and sensory structures of annelids belonging to the Platynereis sp. species complex sampled along the European coast. Using single-cell transcriptomic approaches, this project aims to characterize the cellular changes that help these animals adapt to very different environmental conditions.
The LIFE BRIDGES project, led at the Bork lab (Juery et al.), aims to investigate the molecular basis of associations between bacterial and eukaryotic cells in different biomes and different perturbation regimes. Focusing on genes involved in nutrient exchange, membrane transport, adhesion, and cell–cell communication, we analyse eukaryotic and prokaryotic annotated metagenomes from the TREC expedition to compare human-impacted and pristine environments and evaluate how anthropogenic pressures alter these interaction capacities.

Picoalgae, photosynthetic cells with a diameter smaller than 3 µm, are tiny yet mighty primary producers in many marine oligotrophic environments. The GENOPHY team (Gwenael Piganeau, CNRS) with support from the Roscoff Culture Collection (Ian Probert, CNRS) is establishing the RBM3 picoalgae culture collection using samples from TARA and TREC collected at service sites. This RBM3 collection will provide a valuable resource for the experimental validation of the silico analyses conducted with the VV-club protocols, which explore host-virus interactions. It will also support the generation of high-quality genome, metabolome, and transcriptome reference datasets for cultivable strains, -complementing environmental meta-omics data generated by TREC.

As part of the TREC expedition, the Vincent lab at EMBL investigated how pesticides affect marine microbial communities along the Atlantic coasts of France and Spain. We tested the effects of four widely used pesticides on microbial communities collected from six locations between Lorient and Cadix. Our results will show if and how these four pesticides affect diversity of microbial communities, especially photosynthetic microorganisms.
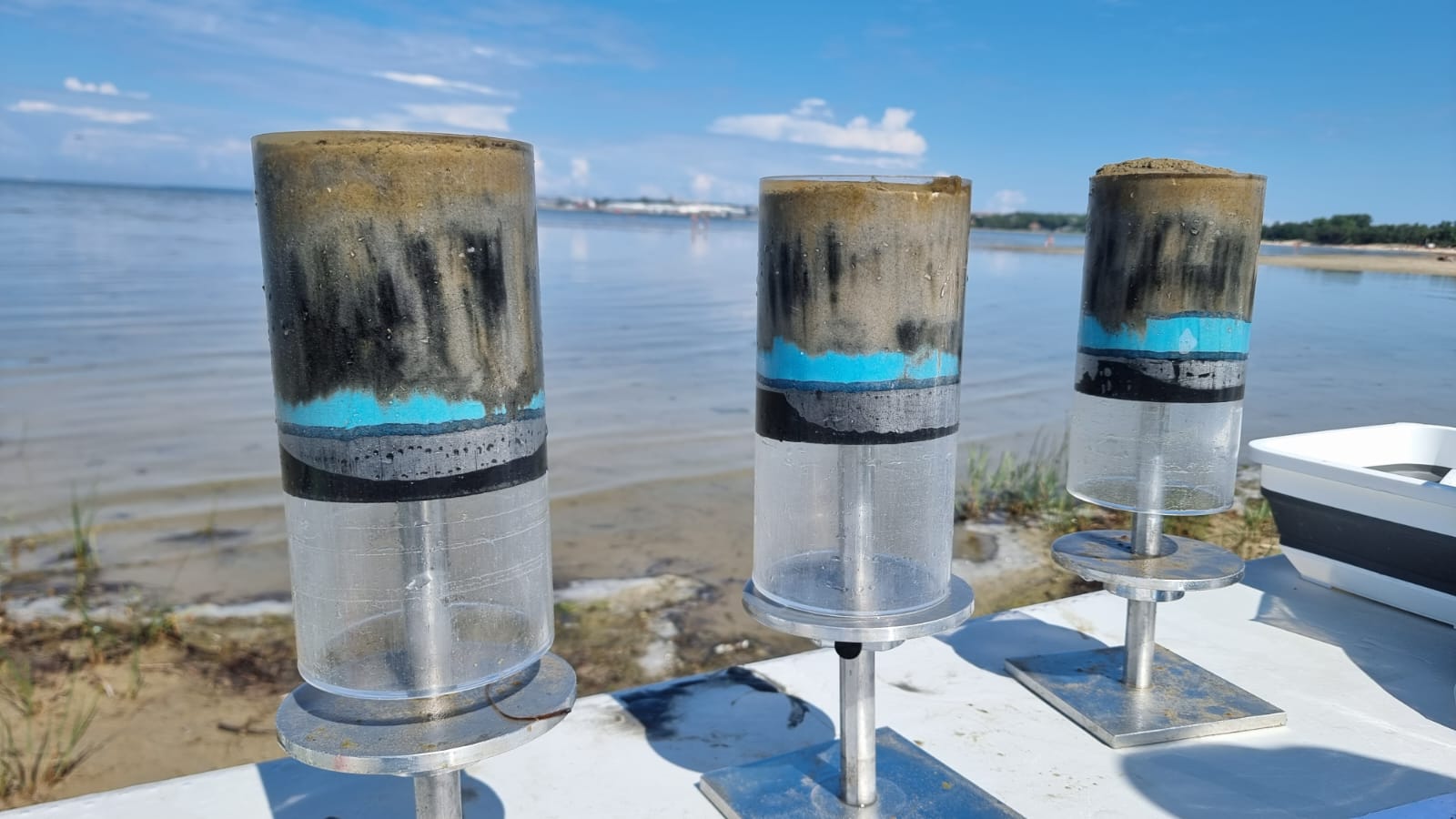

Ifremer and EMBL teams will identify microorganism and meiofauna communities associable to specific chemical pollutions in superficial sediments collected across Europe. This project will assess how chemical contaminants affect taxonomic and functional diversity, and will identify potential indicator species as proxies of chemical contaminations (BIOcean5D and CONTRAST).

As part of the CONTRAST project, two types of optimised methods will be applied to selected TREC sediment samples to map Contaminants of Emerging Concern (CECs). Three types of chemical contaminants (metals and rare earth elements, targeted organic pollutants, and untargeted organic pollutants) will be assessed in surface sediment samples using multi-component analytical tools e.g. TQ-ICP-MS, LC-HRMS, Maldi-TOF. CONTRAST is a Horizon Europe project funded by the EU.
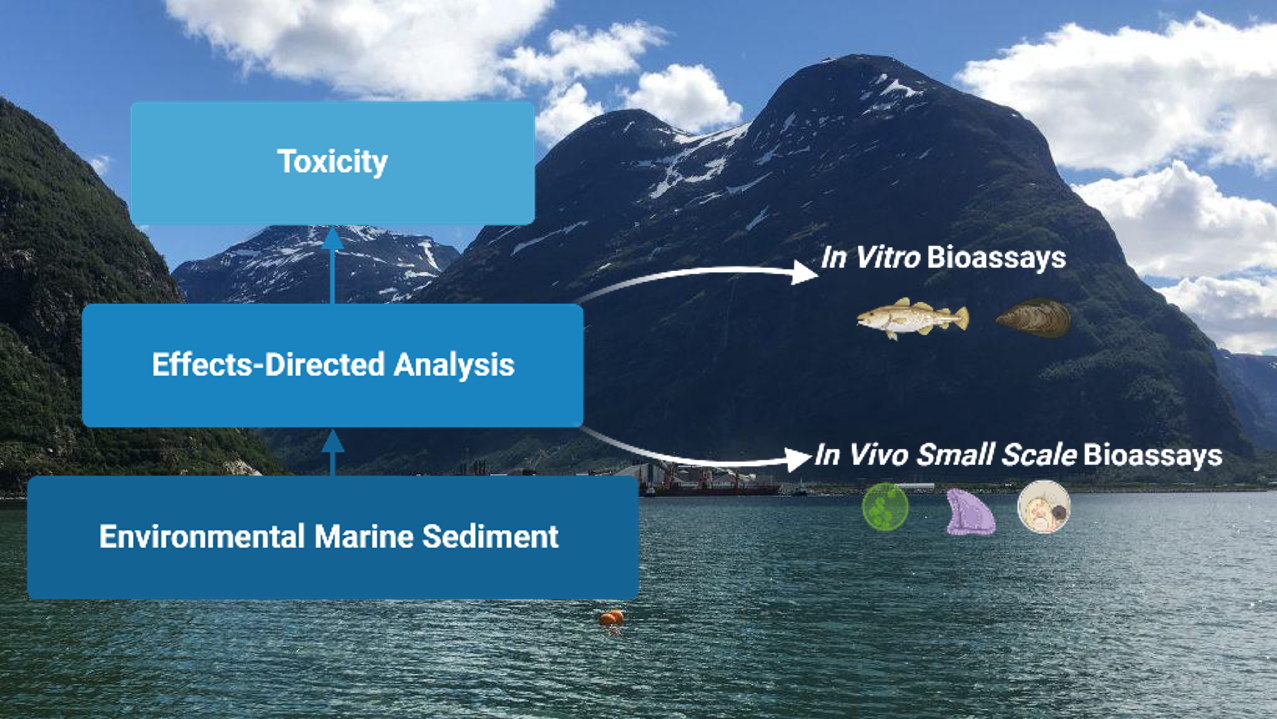
NIVA, IEO, and Ifremer will evaluate the effects of sediment contamination on organisms (microalgae, echinoderms, bivalves, and fish) at different trophic levels and life stages through Effect-Directed Analysis (EDA). Sediments collected along the Mediterranean coastline contain pristine-like areas, areas influenced by agriculture, and those impacted by urbanization and port activities. This work is part of the CONTRAST project, a Horizon Europe initiative funded by the EU.

We aim to utilise barium (Ba) isotopic signatures as a powerful tool for tracing various environmental processes, particularly in coastal ecosystems. By analysing Ba isotopic signatures in soils, rivers, and seawater, we can gain insights into Ba cycling and its interaction with physical and biogeochemical processes. Ba isotopes can help trace long-term changes in river discharges, continental margin inputs, and past ocean productivity. This work will be done at LEMAR (FRA).

This project investigates organophosphate esters (OPEs) and perfluoroalkyl substances (PFAS) in European coastal waters. By combining targeted and non-targeted chemical analysis with microbial data from the TREC expedition, the Vila-Costa group at the IDAEA-CSIC in Barcelona aims to assess pollution sources, distribution, and potential biodegradation of these persistent and toxic compounds.
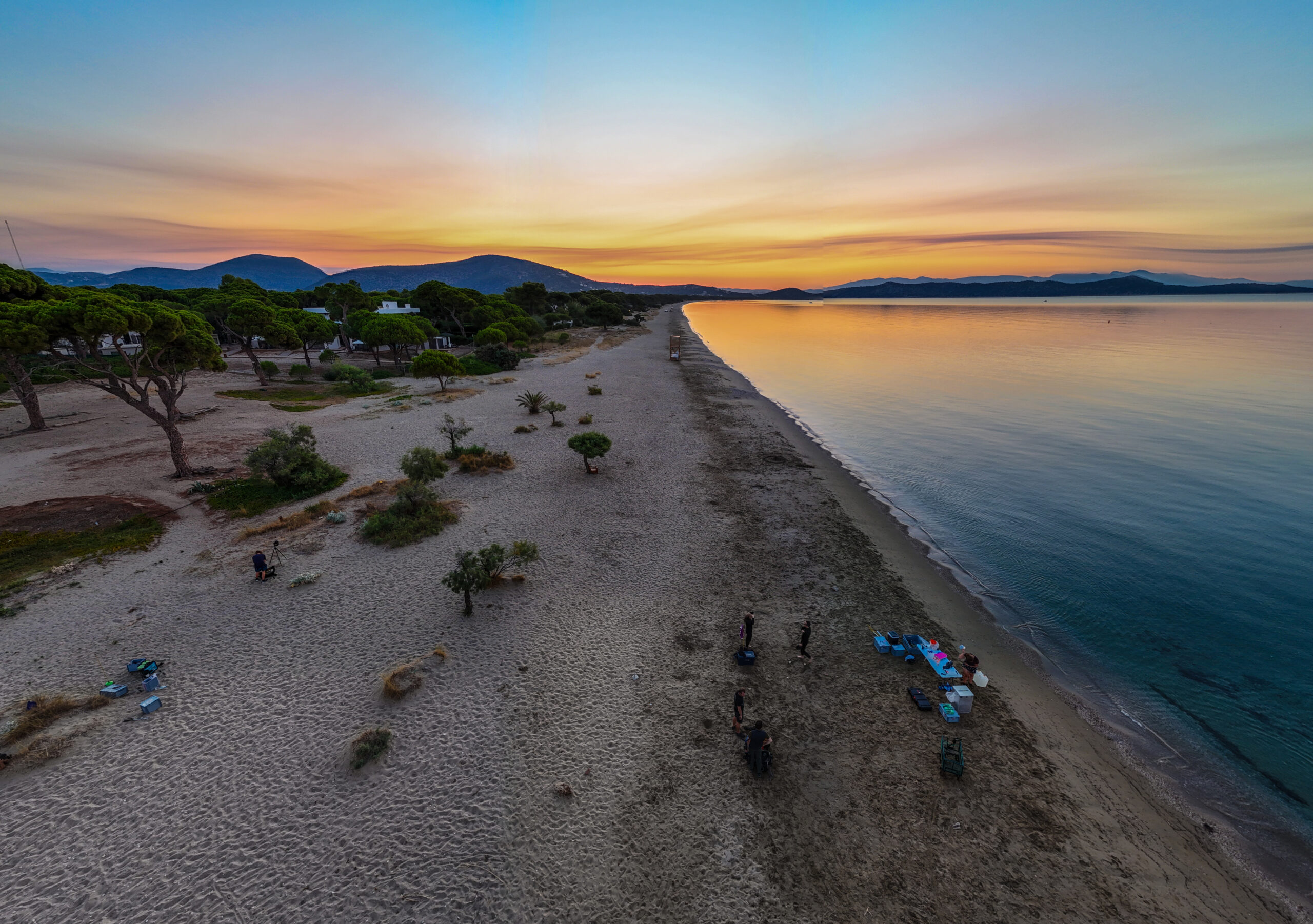

This project aims to study the distribution and the concentration of metals along the European coastline by analyzing a large number of soils and sediments. In addition to the metal distribution, our group (Ifremer and Nantes University) studies the potential sources (natural and anthropogenic sources) and transport of these pollutants. We will also assess the potential risk to wildlife and biodiversity.

Trace metals play a key role in marine ecosystems. Their spatial distribution along Europe’s coasts is essential for assessing ecosystem health and managing human-induced pressures. Originating from both natural and anthropogenic sources, they influence biogeochemical processes and enable the tracing of water masses and ocean dynamics. Their monitoring, led by LEMAR lab (CNRS), contributes to a better understanding of long-term environmental changes, and supports the development of policies to protect coastal zones and adapt to climate change.
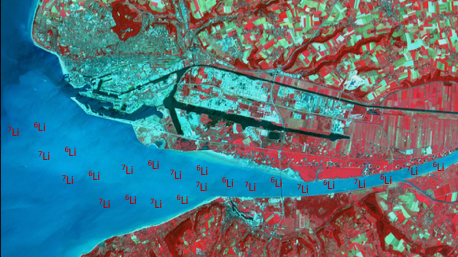

Concentrations and isotopic compositions of lithium will be measured in waters in order to quantify anthropogenic contamination in littoral waters impacted by variable river inputs and anthropogenic activities (LOV, CNR, and supported by ERC Adv SeaLi2Bio project).
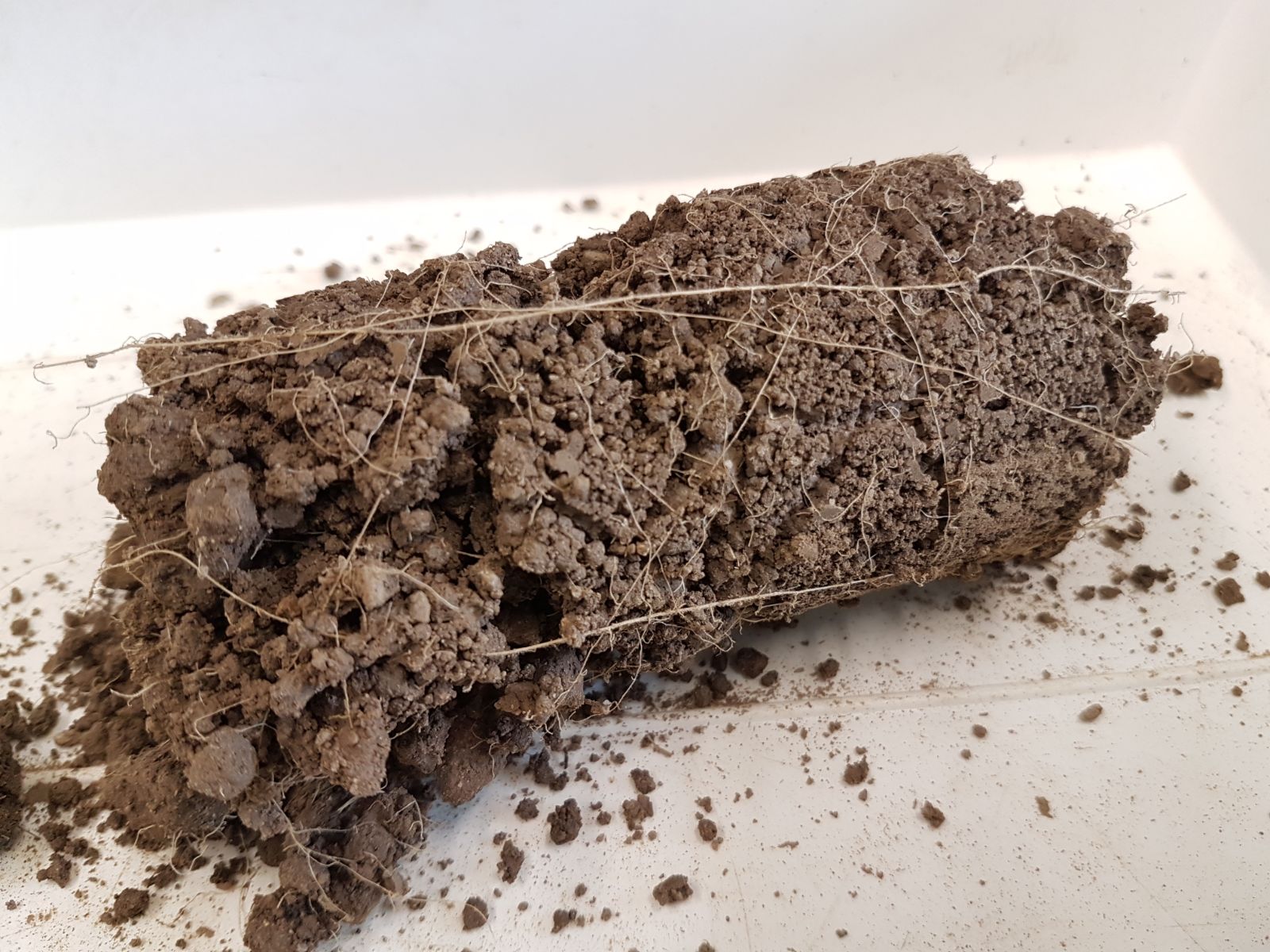

The biogeochemical cycles are strongly related to the living organisms of the ecosystems, from the photosynthesis to the decomposition and persistence functions. Dr Abiven’s team from CNRS and Ecole Normale Supérieure Ulm, will take advantage of the TREC expedition to study carbon, nitrogen and phosphorus cycles in soils, sediments and water, and relate their dynamics and long term persistence to the living organisms.
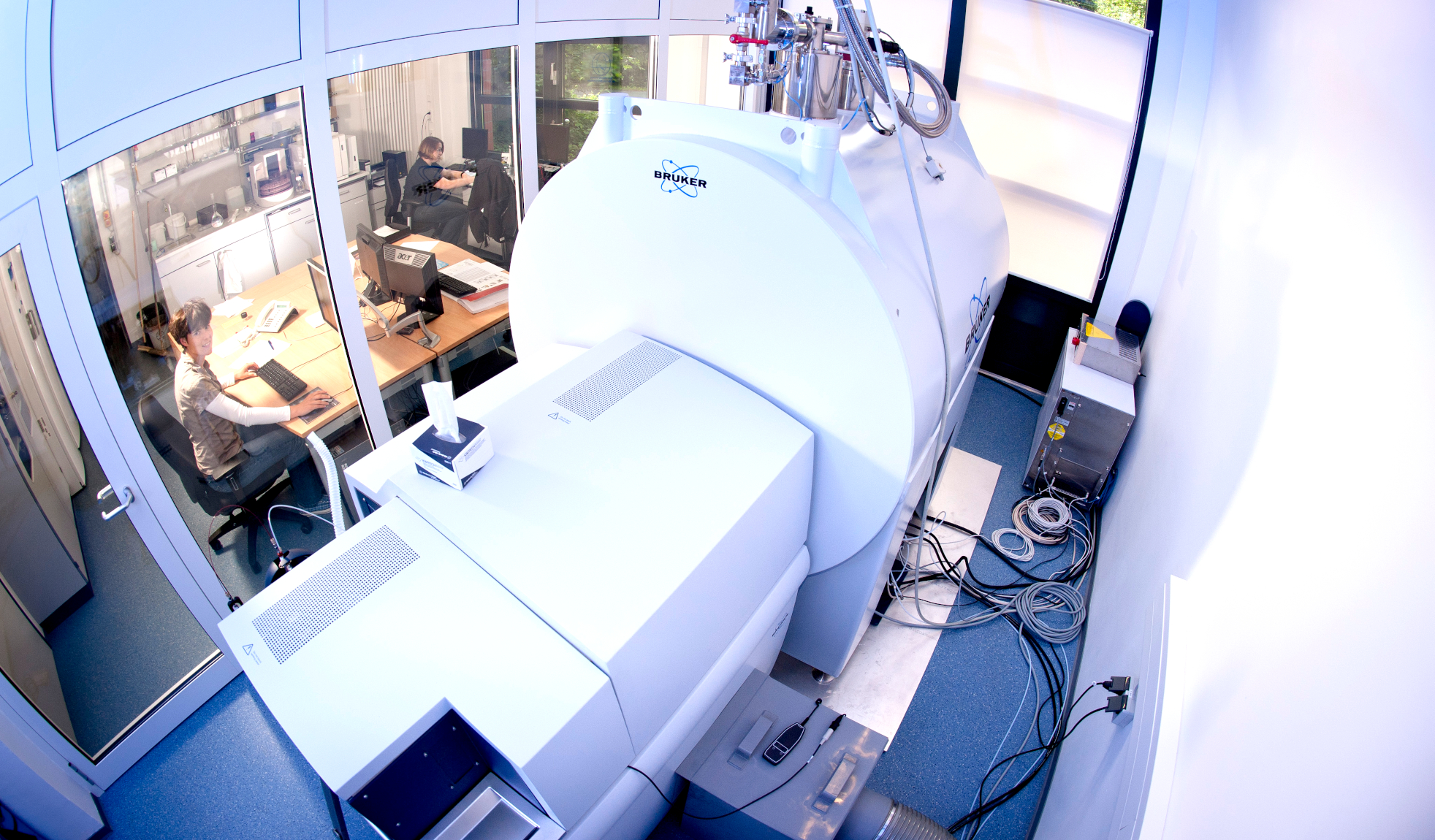

Environmental factors control Microbe-DOM (dissolved organic matter) interactions that result in the consumption, modification and extreme diversification of dissolved organic compounds and are central to marine carbon cycling and sequestration. The interrelations between environmental gradients along the European coast and the fate of DOM will be the focus of Thorsten Dittmar’s lab at the University of Oldenburg during the TREC expedition.
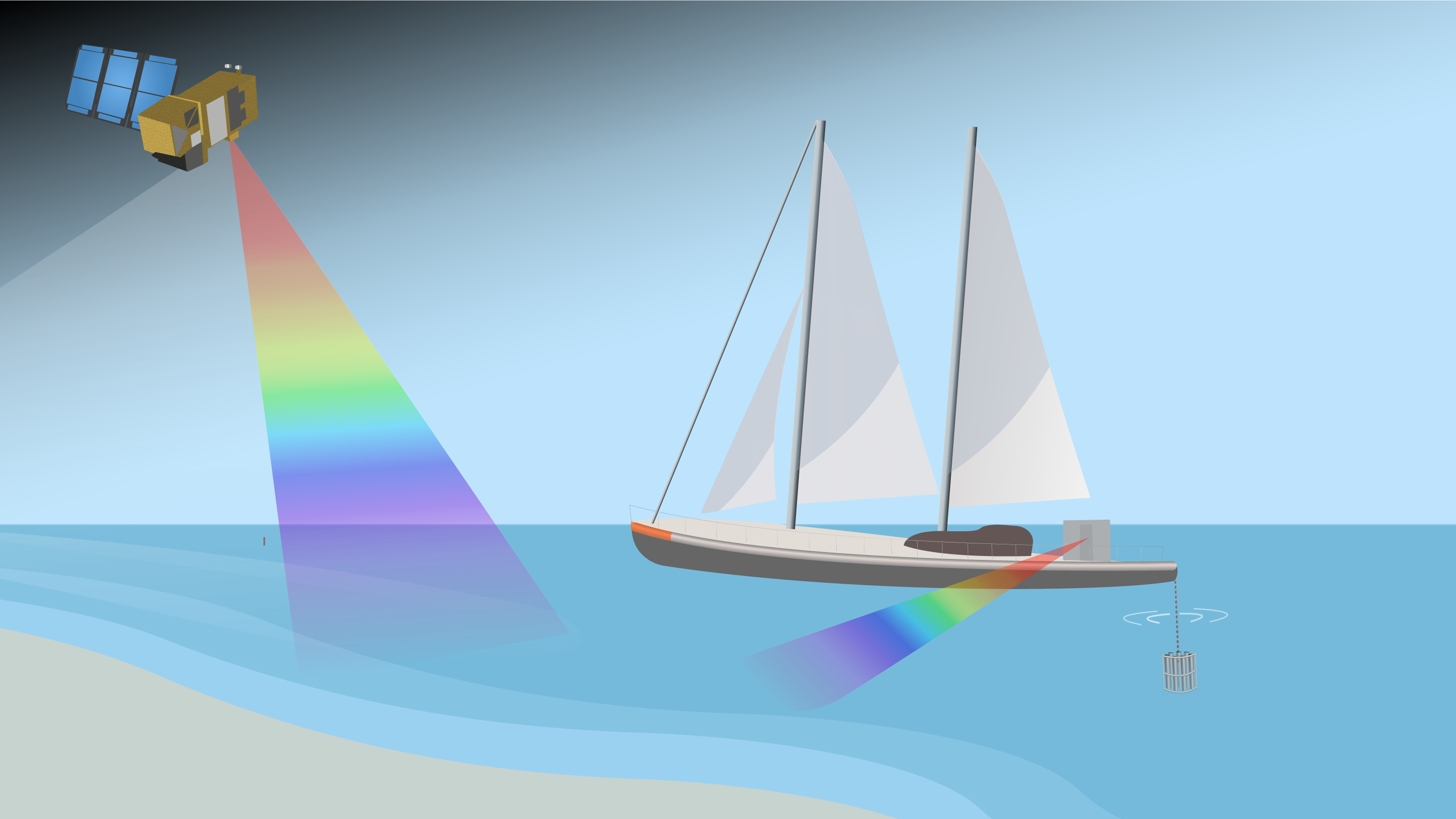

The 2-year project funded by The European Space Agency focuses on the validation of satellite data through in situ observations collected via the TREC (TRaversing European Coastlines) expedition around the European coast.
Using new and existing instrumentation and sampling methods, the project provides vital data for optically complex coastal waters to help monitor biodiversity.
The Zimmermann group (EMBL) will track the abundance of xenobiotics at different sites, focusing on compounds such as pesticides, pharmaceuticals, antibiotics and synthetic hormones. A major aim is to define environmental biomarkers of pollutant exposure, such as microbial taxa or enzymes responsible for degrading these chemicals, or the downstream transformation products released into the environment following microbial degradation.
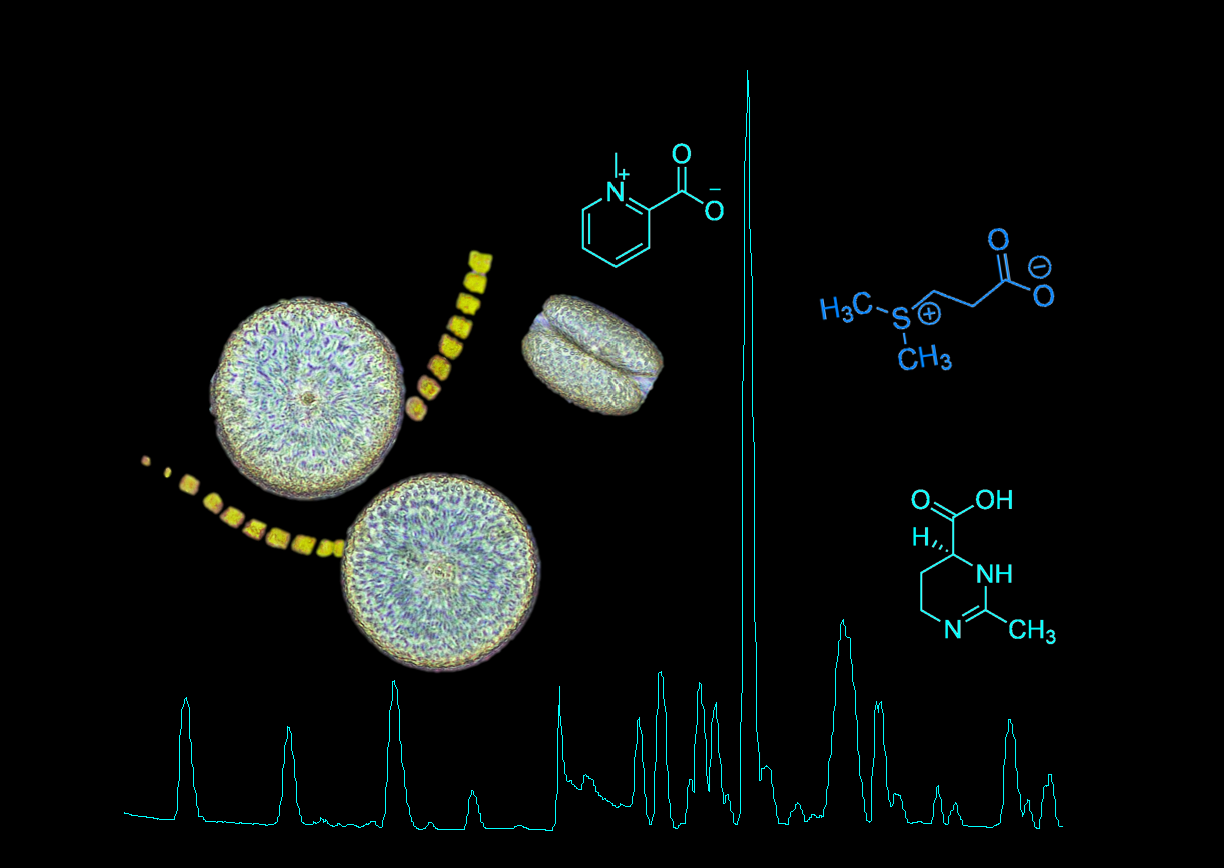
The work of the Pohnert-group focuses on the study of chemical signals within plankton and their impact at the scale of planktonic communities. During the TREC project, we will use Liquid Chromatography coupled with Mass Spectrometry (LC-MS) to determine the chemical fingerprint of marine plankton and elucidate the impact of different environmental factors on the metabolome of the marine microbiome.
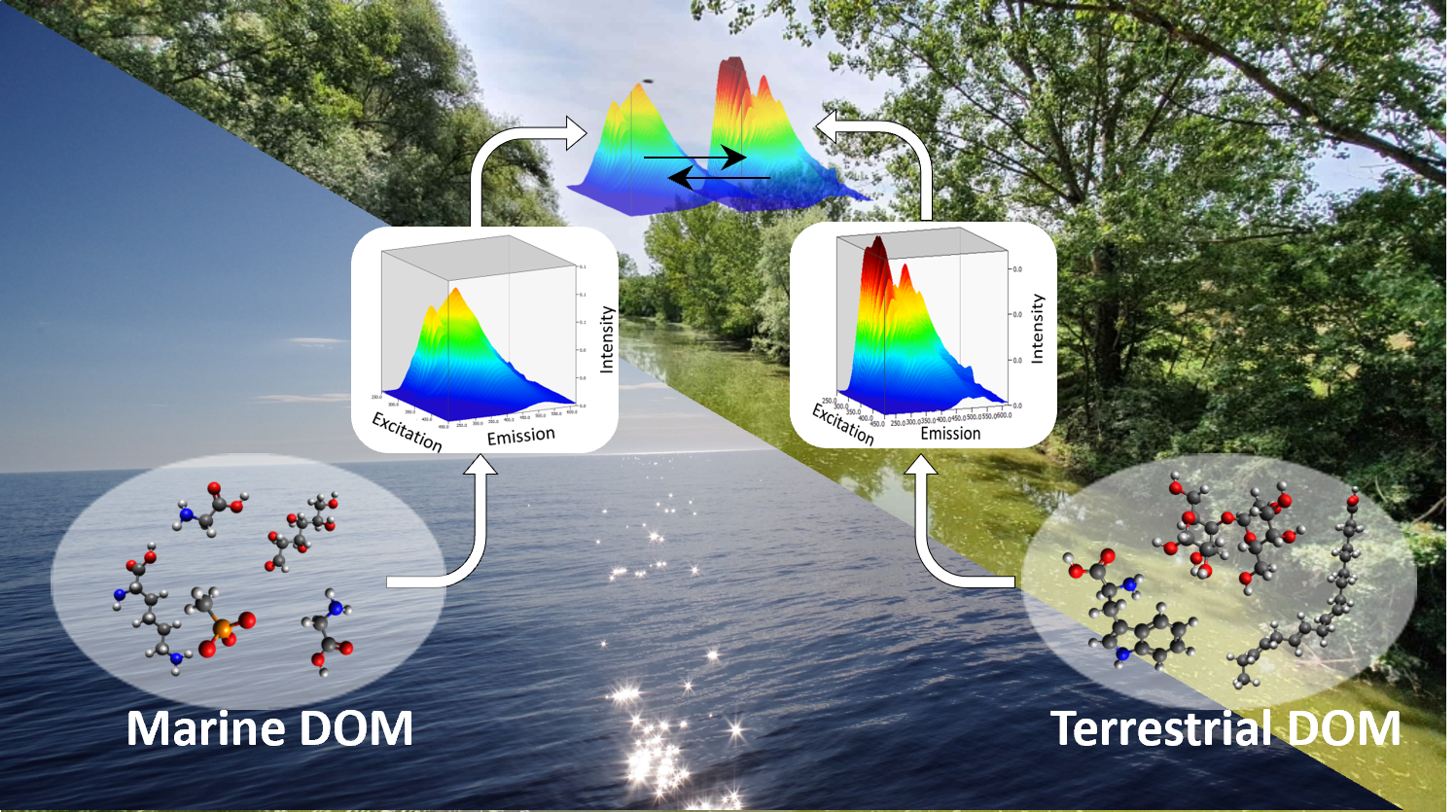
Santinelli’s group at the Biophysics Institute (CNR, Pisa) will gain new insights into the main processes affecting the largest reservoir of organic carbon on our planet (Dissolved Organic Matter, DOM) at the land-sea interface and will quantify the dissolved organic carbon input from land to the coastal European areas. DOM concentration and quality have been reported to affect microbial biodiversity and ecosystem functioning.

This project leverages metagenomics to capture the distribution and dynamics of microbial nutrient cycling pathways. By integrating large datasets from diverse ecosystems, it aims to unveil global patterns of carbon sequestration and nitrogen fixation, highlighting the impact of anthropogenic activities. Led by the Bork group, this research endeavors to provide cross-environment insights into the microbial underpinnings of nutrient cycling.
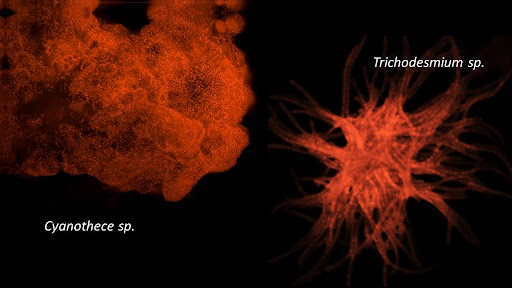
Nitrogen is a key building block for life but its availability is often limited in the ocean. Diazotrophs can overcome this limitation by converting the N2 gas into bioavailable nitrogen. In this project, a team of researchers from Ifremer, CEA, UBO and SZN will measure N2 fixation rates along the mediterranean coastline and combine it with immunolabeling to identify the key players.

This project, led by Ulisse Cardini (SZN), investigates dissolved oxygen, nitrogen, and dimethylsulfide (DMS) saturation in coastal waters to assess key biological processes and understand their role in climate regulation at the land-sea interface. Our work contributes to the Tara Europa and BiOcean5D projects, integrating with TREC to improve climate models and ocean health predictions.
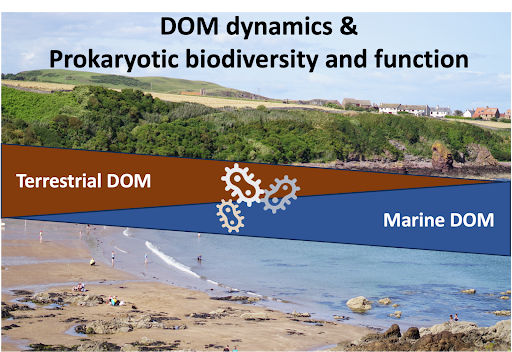
This project investigates the link between microbial diversity and functions and DOM (DOC concentration and optical properties) at the land-sea interface. In collaboration with the Santinelli group in Pisa, the Bork group at EMBL will investigate the role of different pools of DOM (defined based on their optical properties) in shaping the biodiversity and ecosystem functions of marine microbes. This research will enhance our understanding of carbon fluxes and transformations in coastal ecosystems.
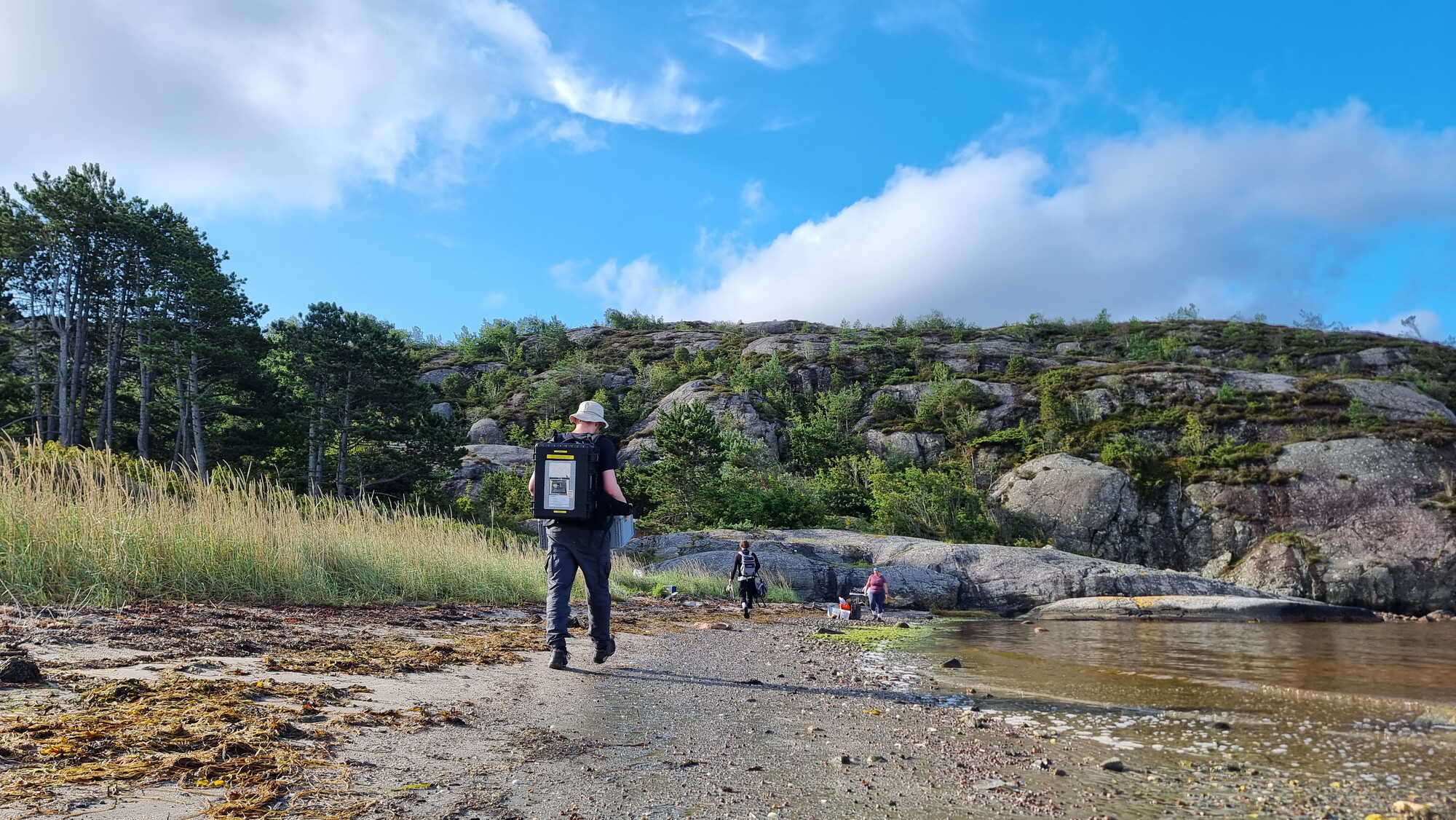

This project will use metagenomic and environmental data from the TREC expedition sampling to study microbial nitrogen cycling. By linking gene abundances and genomic data with pollution and environmental metadata, the Bork group at EMBL aims to understand how human activity shapes the coastal nitrogen cycle and identify novel nitrogen fixers.

This project investigates how human activities, such as pollution and climate change, disrupt microbial nitrogen cycling in oceans and soils. By scaling from TREC samples to global metagenomic data sets, it aims to map nitrogen-related microbial processes, assess pollution effects, and identify bioindicators. Led by the Bork group, this research supports sustainable ecosystem management and pollution monitoring.
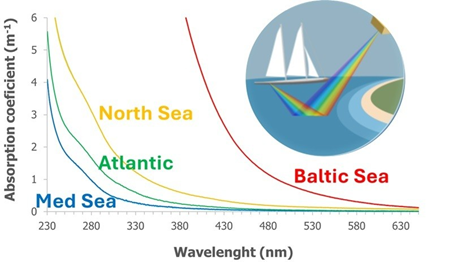
Satellite is a powerful tool to investigate DOM dynamics in coastal areas, where terrestrial inputs vary in both space and time and play a crucial role in carbon budget. With the support of ESA HyperBOOST project, Santinelli’s group (CNR, Pisa), in strict collaboration with Emmanuel Boss (UMaine), will use TREC DOM data to validate available satellite algorithms for CDOM and DOC and to eventually improve their performance, opening to the possibility to fill many of the gaps in knowledge with a focus on anthropogenic activities and climate change impacts on DOM dynamics.
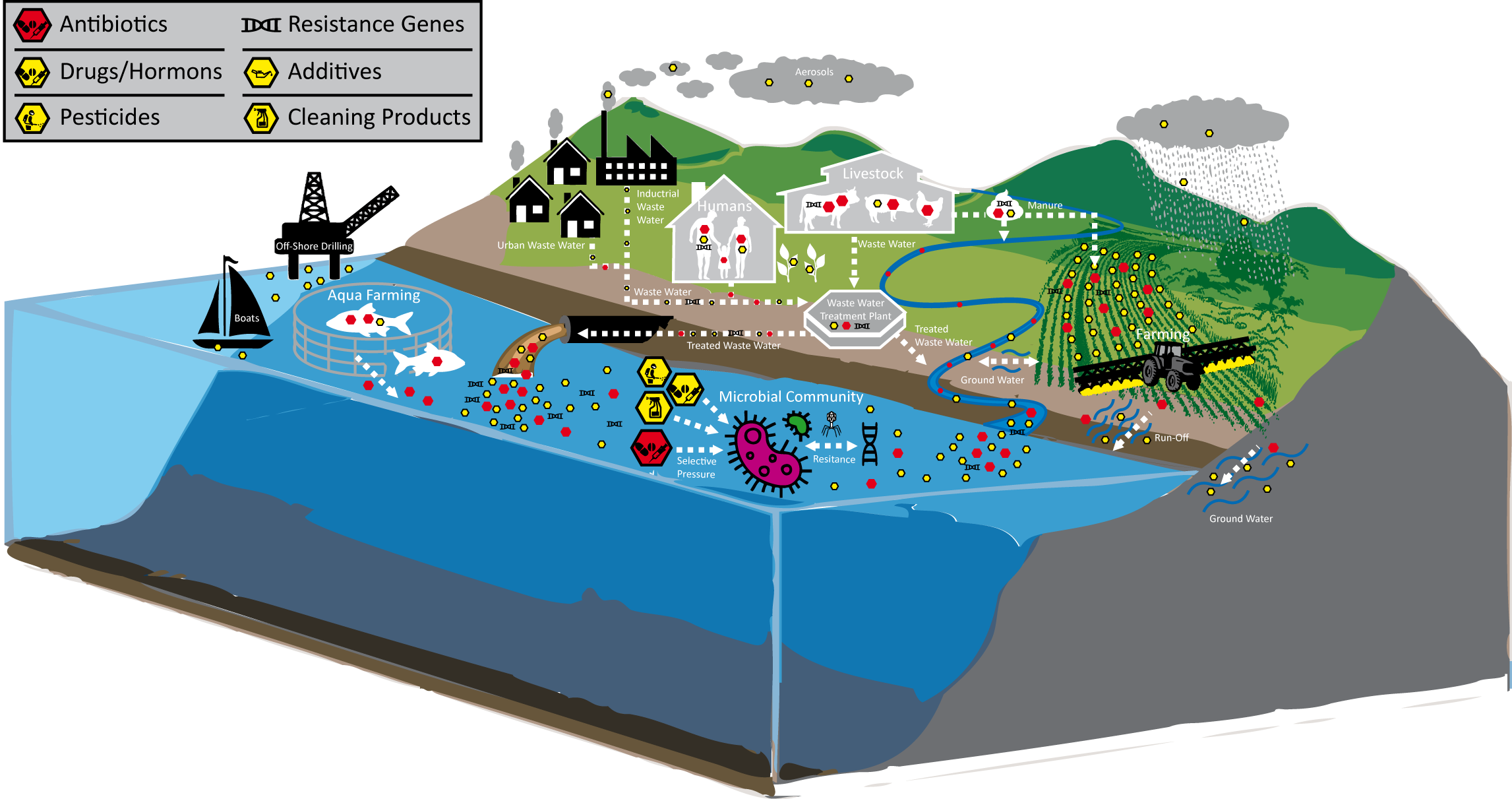
The Petras Group at the University of California Riverside, jointly with the Hass Group at the NIOZ Institute, investigates dissolved organic matter (DOM) and its relation to the marine microbial community. Using non-targeted LC-MS/MS data from TREC samples, we will correlate xenobiotics to antimicrobial resistance-gene abundance and determine how alterations in the DOM chemistry drive microbial community composition and functioning.
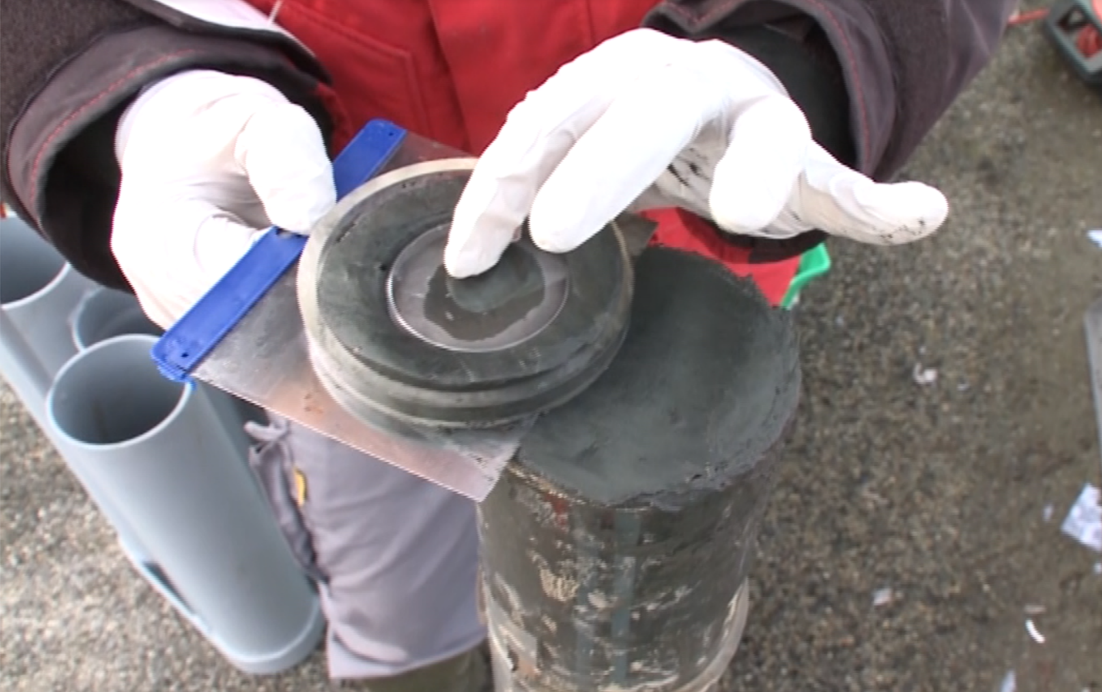

In the frame of the EU BIOcean5D project, Dr Siano (Ifremer) will lead a multidisciplinary research team to assess the effect of human pollution across European coastal ecosystems over the Anthropocene period. Using sediment core bio-geochemical archives, human pollution traces will be associated with inter-site community changes, invasive species dynamics, and emergent risk for human and coastal resources.
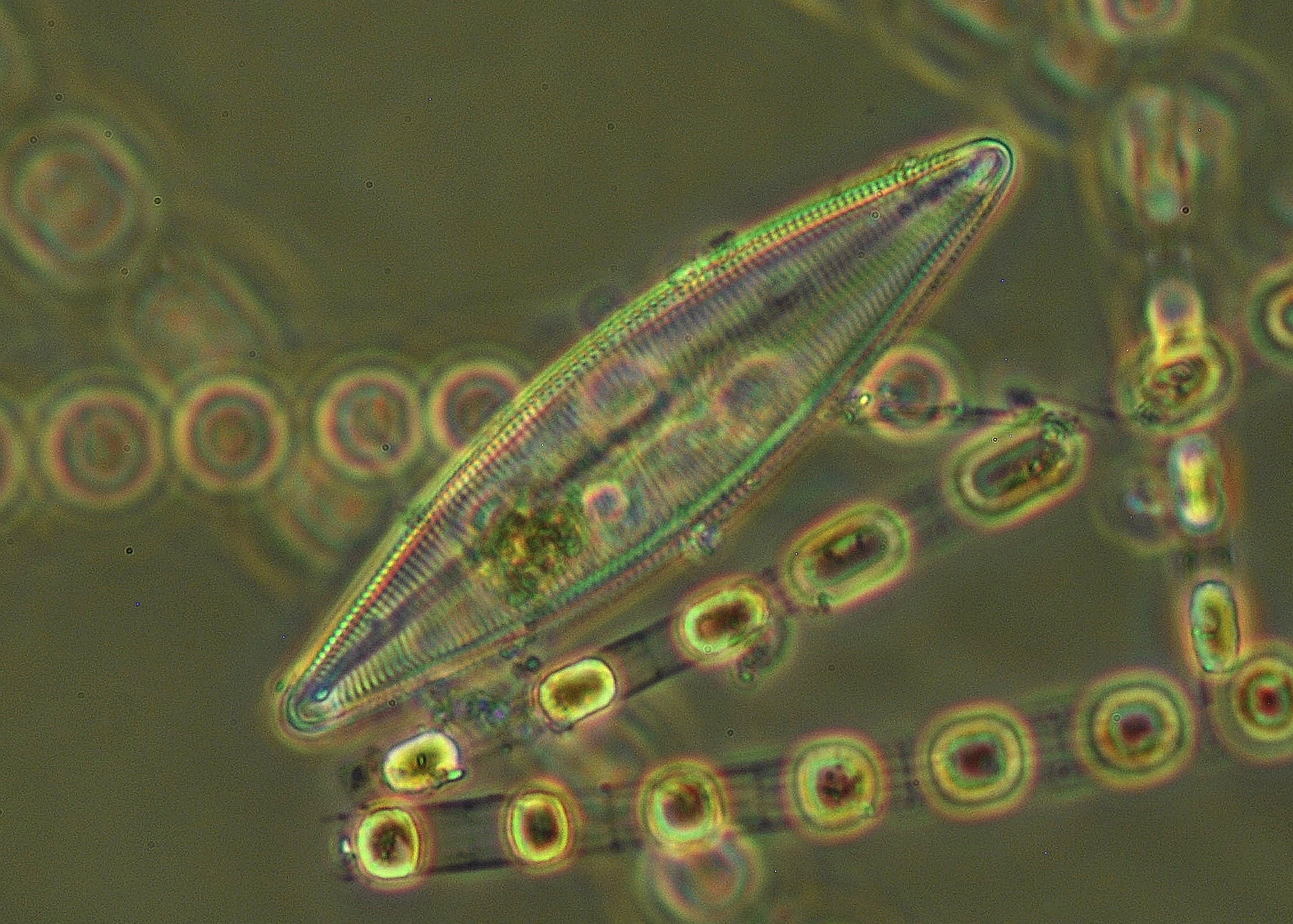
The Sjöqvist group at Turku University will document potential evolution in marine phytoplankton across European waters to clarify the adaptation potential in key species to ongoing climate change. The field of resurrection ecology offers a “backward-in-time” approach via hatching of dormant life stages. This enables the direct quantification of phenotypes and comparison of genetic material across temporal scales.
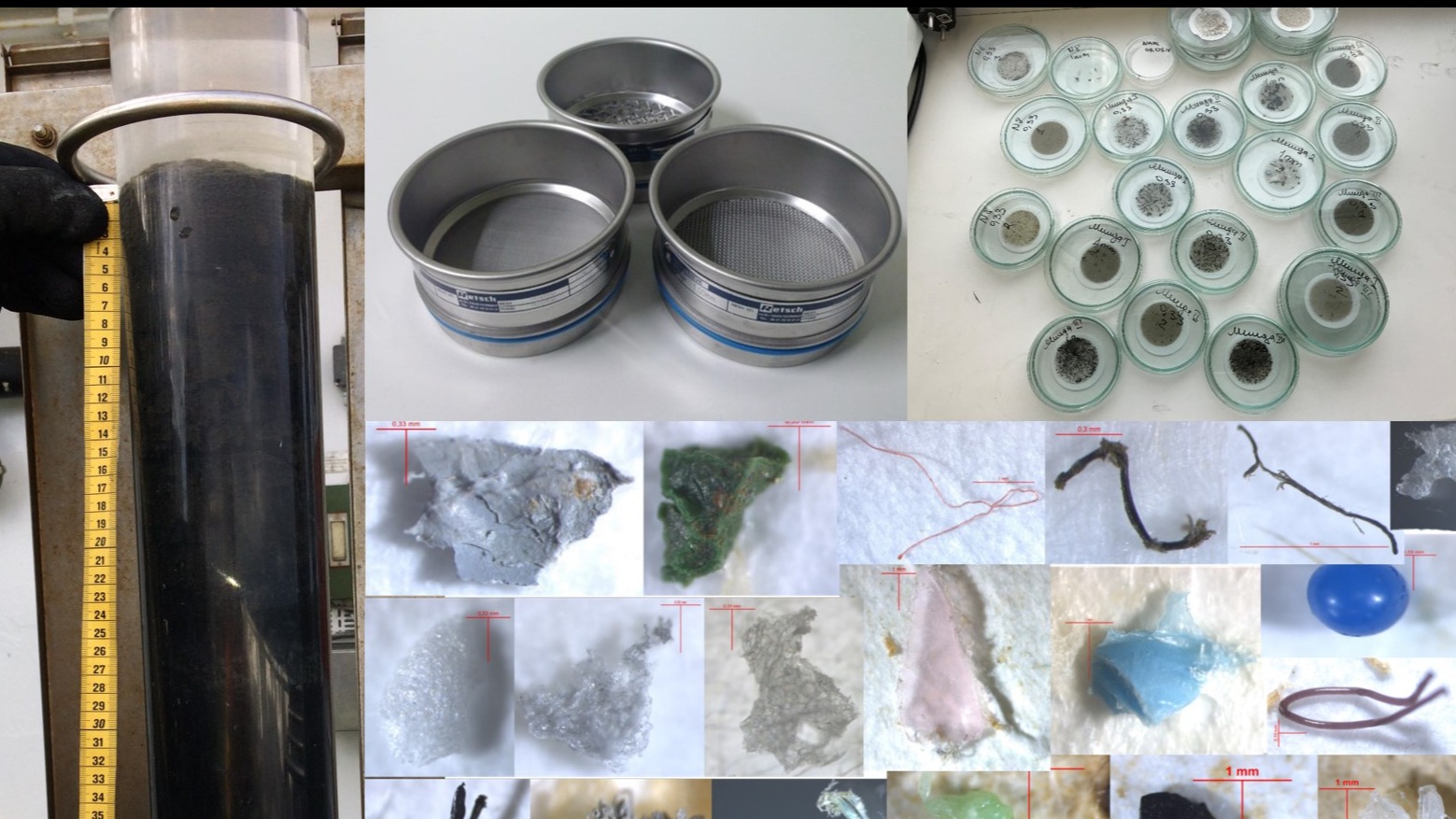

Human-induced eutrophication has a profound effect on the functioning of marine ecosystems. TalTech team investigates historical anthropogenic pollution and its influence on biodiversity and the environmental status of marine ecosystems in different European marine areas. This project focuses on sediment phosphorus storage and release, as well as on deoxygenation of the seabed and bottom water layer.
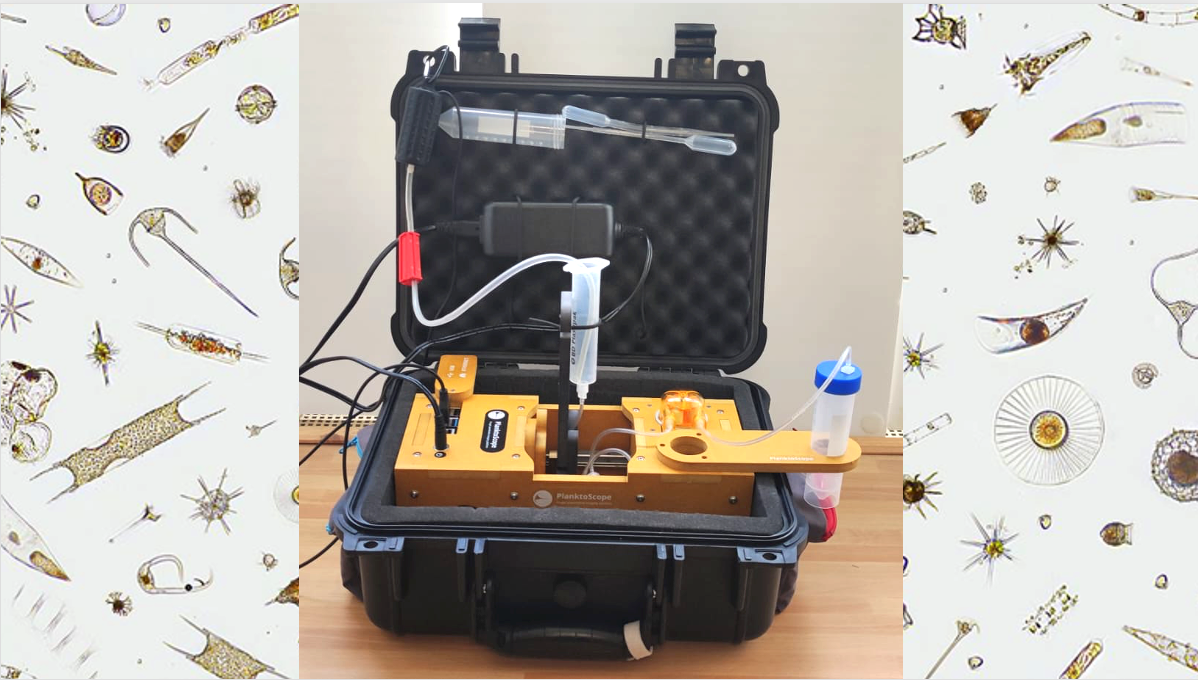

De Vargas and Lombard (CNRS & SU) will deploy innovative frugal tools in research and sea-worker/user communities along the TREC voyage, toward a cooperative and long-term measure of aquatic biodiversity. ‘Seatizen’ will be trained to use the ‘Planktoscope’ and ‘Lamprey’ systems, and generate consistent quantitative imaging and molecular data across ecologically and economically relevant regions, integrated into global databases. (BIOcean5D)

Microbiome science reveals how microbes connect human and environmental health, blurring the boundaries between bodies and their surroundings. The ERC project HealthXCross at the Ca’ Foscari University of Venice, led by Roberta Raffaetà is an ethnographic study examining how interdisciplinary scientific platforms using microbial data across time, space, and species are reshaping health concepts. The project investigates how approaching environments as bodies—and vice versa—transforms: biological diversity understanding, innovation across regions, and health governance.