
Kristina Djinovic Carugo
Head of EMBL Grenoble
ORCID: 0000-0003-0252-2972
EditStructure and assembly mechanisms of sarcomeric cytoskeleton

Head of EMBL Grenoble
ORCID: 0000-0003-0252-2972
EditMovement is vital to all living organisms, from the transport of cellular organelles to the movement of entire organisms. Sarcomeres are the smallest contractile units of striated skeletal and heart muscles. The Z-disc is the attachment region for actin filaments from adjacent sarcomeres and plays a pivotal role not only as an anchoring point of actin filaments, sustaining muscle architecture, but also in signalling, mechanosensing, mechanotransduction, protein turnover, and autophagy. During myofibril formation, Z-bodies, small complexes of assemblies composed of essential Z-discs, grow in size, fuse, and align to form Z-discs (Figure 1). We are interested in the molecular mechanisms underlying the architecture and assembly of Z-discs.
Other research lines focus on studies of bacterial metallo-enzymes involved in protection from chemical and oxidative damage and in catalysing the complete ammonium oxidation cycle.
In order to dissect the Z-disc structure and assembly at a molecular level, we use a combination of biochemical, molecular biophysics, and complementary structural biology methods such as X-ray crystallography, cryo-electron microscopy, small-angle X-ray scattering, atomic force microscopy, and NMR on reconstituted complexes and their individual components. This blend of approaches uncovers biochemical and biophysical properties of macromolecules and their interactions, and gives structural information at different scales of resolution – from atomic to molecular to ultrastructural. These structural and functional data are in turn validated in vitro and in cellula, by structure-informed, molecular, structural, and cellular biophysics studies (the last in collaboration). Finally, we combine these experimental data with integrative modelling to generate structural models of Z-disc assemblies.
In parallel, we are developing methods for rectification of protein generation, crystallogenesis, and for structural studies and modelling of intrinsically disordered proteins (Mlynek et al., Amino Acids, 2014; Mlynek et al., Crystal Growth & Design, 2019; Kotov et al., Prot Sci, 2021).
Structure and PIP2 regulation mechanism of human α-actinin
Actin is one of the most abundant proteins in eukaryotic cells. Actin filaments, together with a large number of actin-binding proteins, are critical players in many cellular functions, ranging from cell motility and muscle contraction to maintenance of cell shape and transcription regulation. α-Actinin, an archetypical F-actin-binding and bundling protein, is the major protein in muscle Z-discs, where it cross-links actin filaments from adjacent sarcomeres. We determined the first complete structure of α-actinin from striated muscle and explored its functional implications on the biochemical and cellular levels. The structure provides insight into the mechanism of α-actinin regulation by PiP2 and the intramolecular pseudoligand autoinhibition mechanism. It also offers a foundation to study interactions with Z-disc partners and effects of pathogenic mutations with molecular resolution and mechanistic insight (Ribeiro et al., Cell, 2014).
Structure and calcium regulation mechanism of ancestral α-actinin
While muscle α-actinin is regulated by PiP2, Ca2+ inhibits α-actinin capacity to bundle actin filaments in non-muscle cells. We determined the structure of an ancestral Ca2+-regulated α-actinin from Entamoeba histolytica and proposed a molecular mechanism for its regulation. We uncovered that Ca2+ binding triggers an increase in protein rigidity, leading to reduced conformational flexibility and bundling activity. The proposed molecular mechanism is likely to be a blueprint for the regulation of spectrin-like proteins (Pinotsis et al., PNAS, 2020).
Myotilin as a regulator of sarcomere assembly
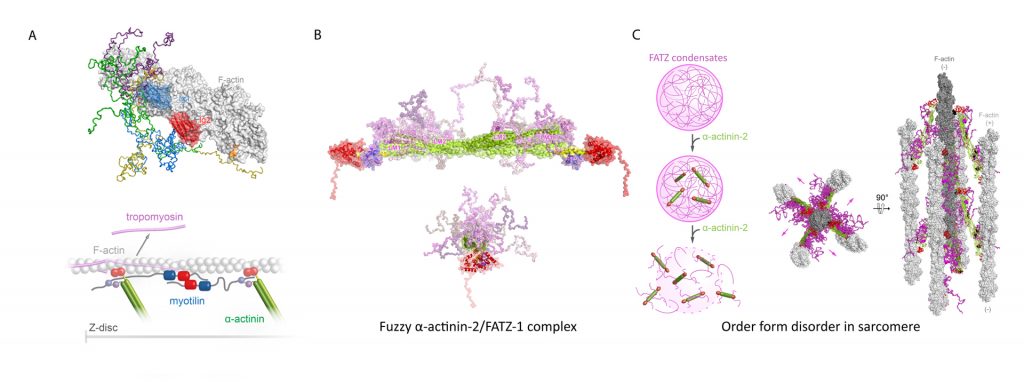
Myotilin is a Z-disc scaffold protein that provides structural integrity by multiple interactions, including with actin filaments and α-actinin-2. We generated the first integrative structural model of its complex with actin filaments. We further showed that myotilin displaces tropomyosin from F-actin, implying a novel role of myotilin in sarcomere biogenesis beyond acting as an interaction hub for Z-disc partners (Figure 2A) (Kostan et al., PLOS Biol, 2021).
Order from disorder in the sarcomere
In sarcomeres, α-actinin crosslinks actin filaments and anchors them to the Z-discs. FATZ proteins interact with α-actinin and five other core Z-disk proteins, contributing to myofibril assembly and maintenance as a protein interaction hub. We determined the first structure of α-actinin-2 in complex with a Z-disk partner, FATZ-1, which is best described as a conformational ensemble, and carried out its cellular validation. We showed that FATZ-1 forms a tight fuzzy complex with α-actinin-2 and proposed a molecular interaction mechanism via main molecular recognition elements and secondary binding sites. The obtained integrative model reveals a polar architecture of the complex which, in combination with FATZ-1 multivalent scaffold function, might organise interaction partners and stabilise α-actinin-2 preferential orientation in the Z-discs (Figure 2B). Furthermore, we uncovered FATZ-1 ability to phase-separate and form biomolecular condensates with α-actinin-2, raising the tantalizing question of whether FATZ proteins can create an interaction hub for Z-disc proteins through membrane-less compartmentalisation during myofibrillogenesis (Figure 2C) (Sponga et al., Sci Adv, 2021).
Our future goals are to contribute to the mechanistic understanding of the following questions: