
James Sharpe
Head of EMBL Barcelona
ORCID: 0000-0002-1434-9743
EditMulticellular systems biology

Head of EMBL Barcelona
ORCID: 0000-0002-1434-9743
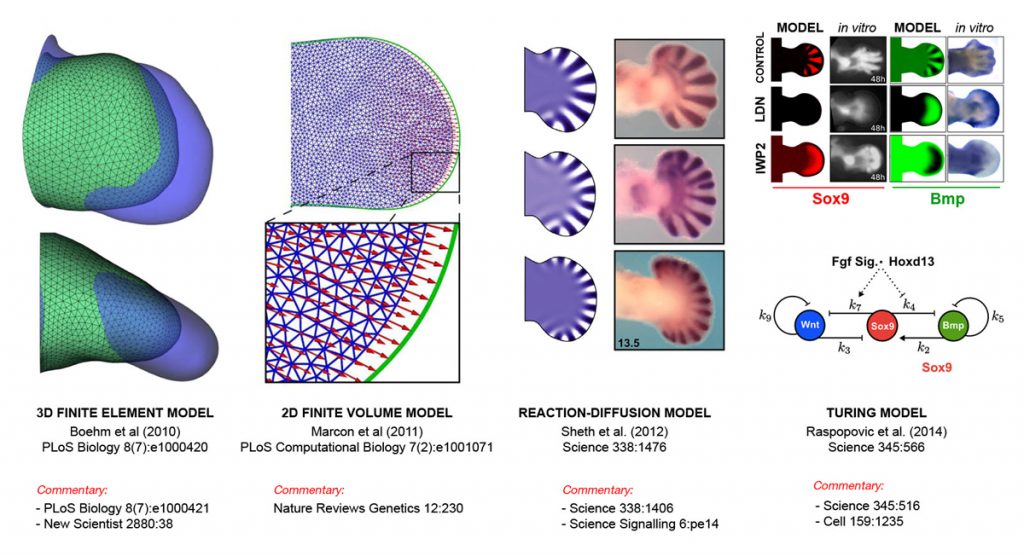
EditHow do networks of genes control populations of cells to build organised tissues and organs? Organ development is a multi-scale process, in which molecular and cellular events control large-scale tissue movements, but these ‘macroscopic’ dynamics are equally important in feeding-back to regulate molecular events. A full understanding of how genes control organogenesis will thus require multi-scale computer modelling, and we have chosen vertebrate limb development as a model system to explore this problem.
Such a model should be based on quantitative data about dynamic tissue shapes and spatial distributions of gene activities. Traditional high-throughput and ‘omics’ technologies do not preserve spatial information, and we have therefore been developing and pioneering various 3D mesoscopic imaging technologies to generate geometric and spatial data for the model.
Our research thus falls primarily into two main areas:

Researchers at EMBL Barcelona have developed an open-source tool that makes working with complicated volumetric imaging data easier.
Edit
LimbNET is a new online platform that integrates computer modelling, experimental data, and 2D live simulations.
Edit
EMBL researchers have been using a powerful technique called ultrastructure expansion microscopy to peek deeper inside living organisms and understand how they function.
Edit
Better than the original fairy tale, Kendrew awardee Irma Querques’s ‘Sleeping Beauty’ story offers potential for new therapeutics.
Edit
Researchers at EMBL Barcelona have quantitatively evaluated the risk of undetected artefacts in single-cell RNA sequencing experiments.
Edit
A recent pan-EMBL event provided an opportunity to reflect on responsible research assessment in scientific institutions.
Edit
As Peer Bork and Ewan Birney take up interim leadership of EMBL, the organisation announces additional changes in site leadership.
Edit
EMBL Barcelona researchers developed a computational method that reconstructs embryonic development.
Edit