Previous and current research
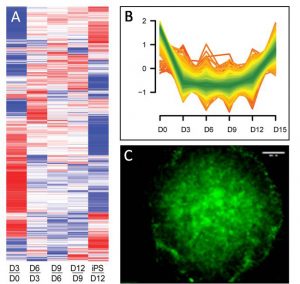
Proteomes are highly complex, consisting of thousands of proteins that operate in intricate networks in a cell and condition-specific manner. Our main interest is to develop and apply mass spectrometry-based approaches to understand how dynamic regulation of the proteome underlies processes that are fundamental to cancer biology and to pluripotency in stem cells. For instance, this has enabled us to identify novel proteins controlling the identity of mouse hematopoietic stem cells, and proteins that are key in the gain of pluripotency during reprogramming of fibroblasts to induced pluripotent stem cells (iPSCs). In addition, we combine this with biochemical techniques allowing us to focus on specific sub-proteomes, to get a deeper insight in the spatial regulation of the proteome when exposing cells to defined perturbation conditions (e.g. stress, drugs or growth factors). This focuses on secreted proteins for their role in cellular crosstalk, in RNA-binding proteins involved in translational control, and in chromatin-binding proteins for their key function in transcriptional regulation and cell fate decision. We have developed dedicated biochemical methodologies to enrich for each of these protein classes, based on click-chemistry to study secretory proteins, and combinations of cross-linking and various affinity-enrichment approaches to capture and identify proteins interacting with RNA and DNA. For instance, we have used this to characterize chromatin-bound protein networks in mouse embryonic stem cells, identifying novel proteins that we showed to functionally contribute to establishing pluripotency.
Future projects and goals
We will exploit the power of these innovative biochemical methods in combination with our mass spectrometric platforms to gain mechanistic insight in the regulation of cellular plasticity in cancer and stem cell identity. In particular, we will use our optimised workflows for global proteome profiling of clinical tumour samples. Secretome analysis will shed light on inter-cellular communication in a cancer context, and on principal mechanisms of secretory pathways. We will extend our chromatin interaction studies to identify and functionally characterize the composition and dynamics of protein networks around transcription factors and chromatin-modifying enzymes. We increasingly combine this information with ChIP-seq and RNA-seq, to gain a more complete understanding of how gene expression is regulated, and how this is derailed in cancer.