
Simone Koehler
Group Leader
ORCID: 0000-0002-1119-8855
EditSelf-organisation in meiosis

Group Leader
ORCID: 0000-0002-1119-8855
EditMeiosis is a specialised cell division process that generates haploid gametes from diploid precursor cells. During the meiotic cell division process, homologous chromosomes undergo recombination and are then segregated from each other. Errors in this process are a major cause of infertility and can also result in congenital conditions such as Down syndrome. For homologous chromosomes to segregate properly during meiotic divisions, they must first pair during meiotic prophase. Pairing is ultimately stabilised by crossover formation, which generates physical linkages between the homologous chromosomes and gives rise to genetic diversity. However, how these processes are regulated remains unclear to this date. More than a century ago, scientists identified a unique, ladder-like structure, the synaptonemal complex, using electron microscopy. The synaptonemal complex assembles between homologous chromosomes during meiosis. Since then, several studies have identified the components within this complex and have revealed that this structure is key in regulating a multitude of meiotic processes.
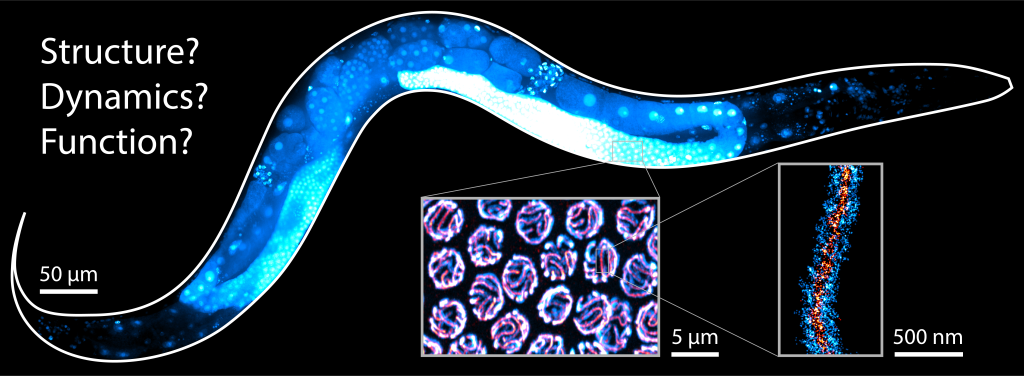
Despite its wide conservation and its central role in meiosis, little is known about the molecular organisation and function of the synaptonemal complex. Our group uses microscopy techniques ranging from super-resolution microscopy to in vivo time lapse imaging in the model organism Caenorhabditis elegans to dissect the ultrastructure and molecular mechanisms of the synaptonemal complex.
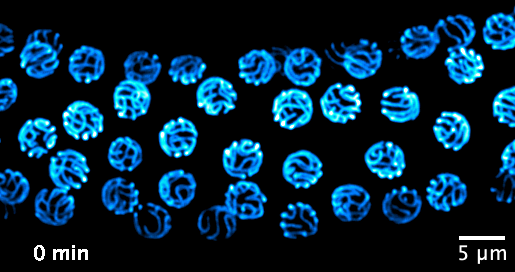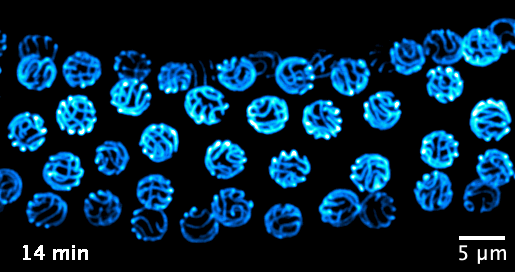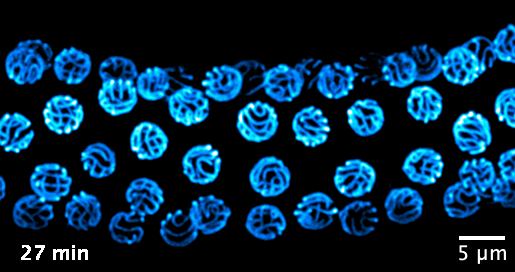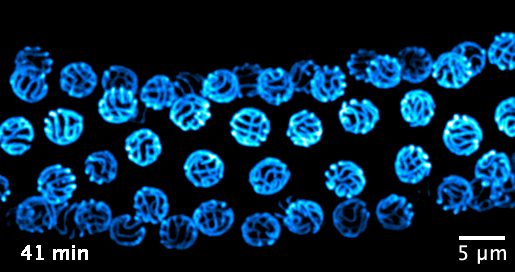
We now focus on the mechanisms that govern the formation and function of the synaptonemal complex, and its role in chromatin organisation. We aim to investigate the following questions:
1. How do homologous chromosomes recognize each other and assemble the synaptonemal complex?
2. How is crossover formation regulated? What is the role of the synaptonemal complex?
3. What are the dynamic processes required to ensure robust and correct pairing, synapsis, and recombination during meiosis?
To address these questions, we take advantage of genome editing, genetics, and biochemistry in combination with state-of-the-art imaging techniques to image meiosis in situ and in vivo. We also develop image analysis tools to obtain quantitative insight into the processes that drive and regulate homologous chromosome pairing, synapsis, and crossover formation.