Previous and current research
Pluripotent cells have the dual ability to self-renew and differentiate. Therefore, in pluripotent cells, the expression of hundreds of genes should be stable in the self-renewal case, but gene expression can also be directed in a coordinated manner towards particular states upon external signalling cues (lineage commitment towards terminal differentiation). Deciphering this complex problem has garnered much attention at the systems level.
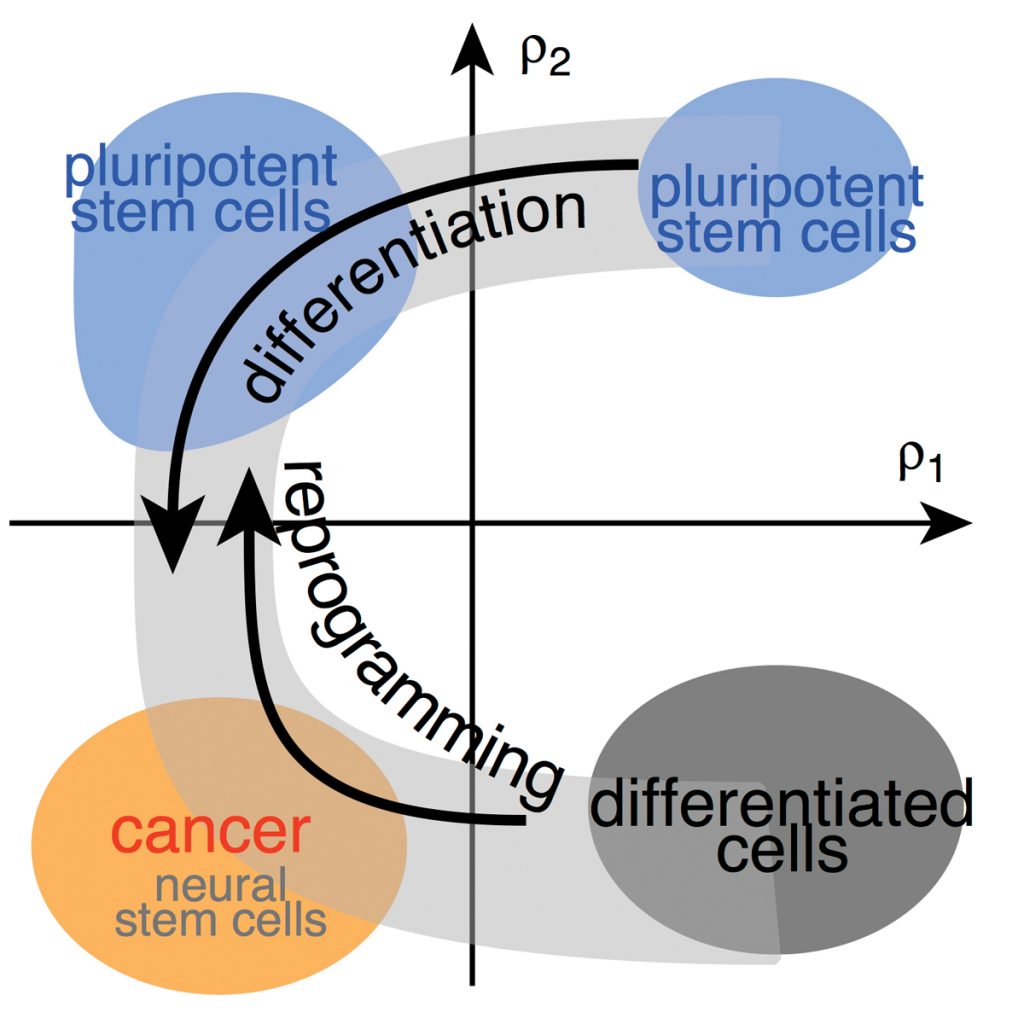
Tackling this challenge requires good characterisation of the pluripotent state. miRNAs are suitable marker candidates because they are excellent classifiers of tissue types or cellular states and they also play a crucial role in differentiation. By profiling miRNA expression in human cells, we have previously shown that pluripotency surprisingly emerges as a much more diverse state than previously believed: variability in miRNA expression is comparable to that found in differentiated cells and cancer cells. We have also shown that it is possible to dramatically reduce the complexity of miRNA expression patterns to a few meaningful dimensions, suggesting that complex processes of the stem cell system, such as differentiation and reprogramming, can be mapped quantitatively.
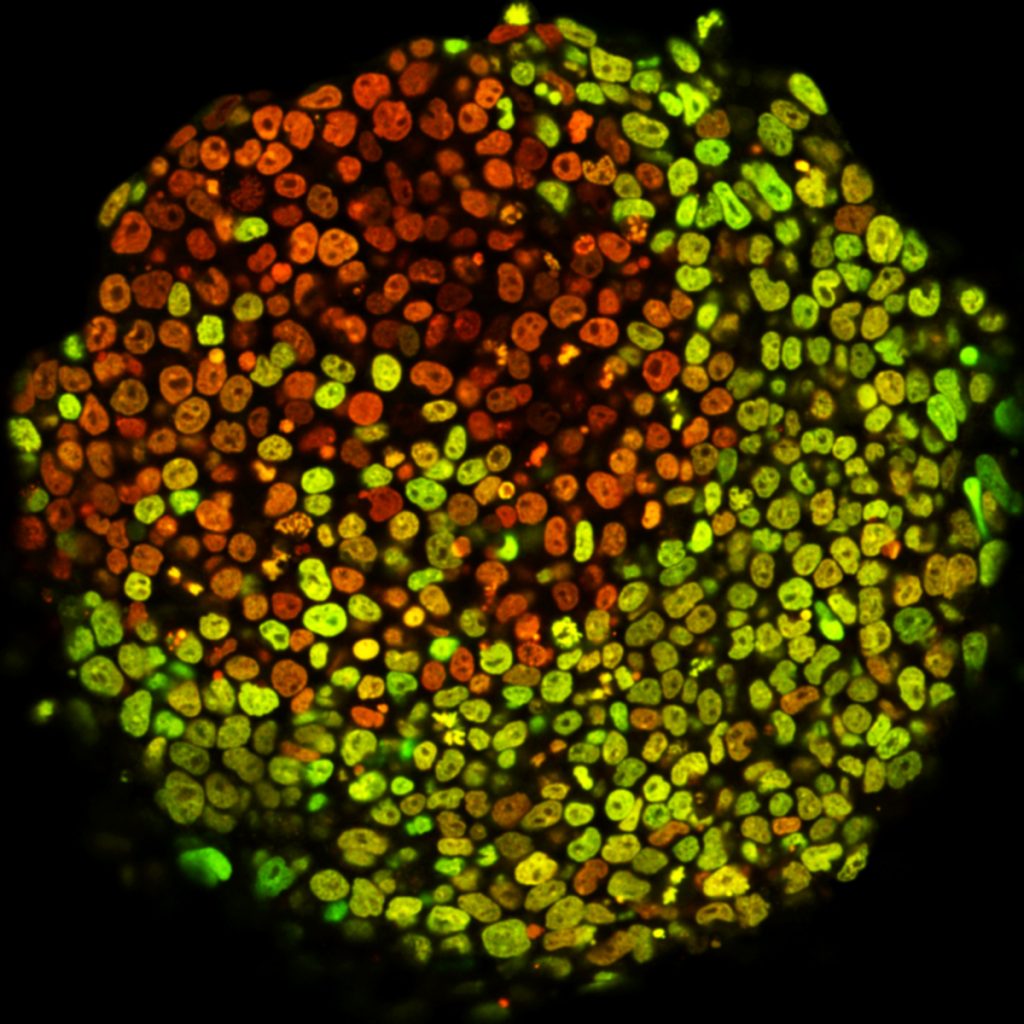
Currently, we are employing a dynamic approach at the single cell level to resolve the dynamics of differentiation and the different molecular and cellular processes at play during fate determination. Indeed, differentiation is intrinsically a dynamic process, where individual cells have to change from one state to another. Having developed fluorescent reporters to assess miRNA expression in single cells, we are characterising mouse embryonic stem cell (ESC) self-renewal using single-cell live imaging. We have found that the miRNA miR-142 is bimodally expressed in self-renewing mouse ESCs. miR-142 expression stratifies ESCs into two distinct subpopulations that interconvert stochastically: one subpopulation able to differentiate and a subpopulation deaf to differentiation cues. miR-142 expression creates this phenotypic diversity by switching the activation status of key intracellular signalling pathways.
Future projects and goals
We plan to study the dynamics of differentiation at the single-cell level both in vitro in mouse embryonic stem cells and in vivo. The ultimate goal is to dissect the transcriptional regulation and gene networks and the associated cellular changes underlying stem cell differentiation. We are taking an integrated systems biology approach that combines single-cell live imaging of miRNA expression, image processing, perturbation approaches, and mathematical modelling. We wish to address the following questions:
- How dynamic is the pluripotent state?
- What are the in vitro dynamics of differentiation of mouse ESCs?
- How do in vitro findings compare to in vivo differentiation behaviour?