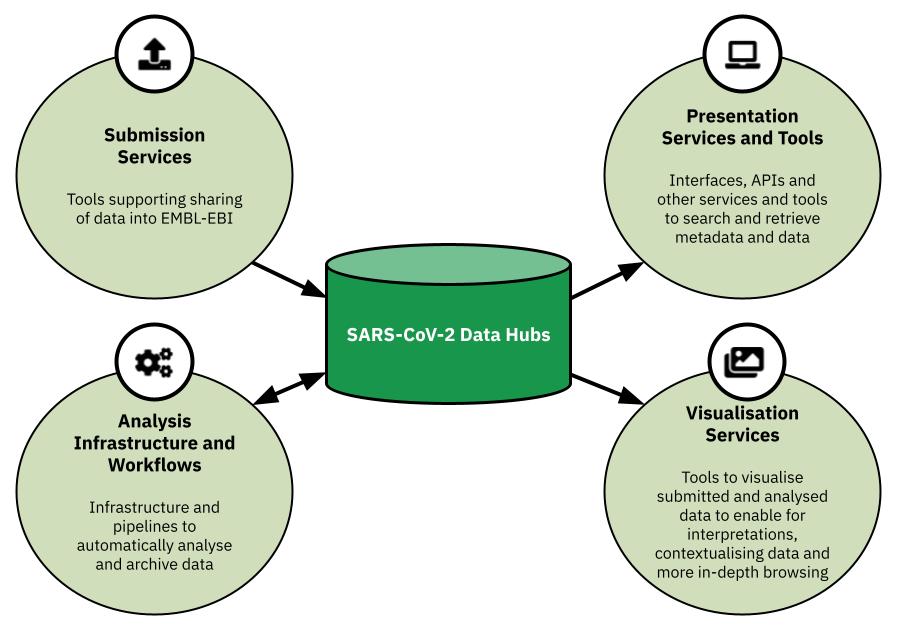
The SARS-CoV-2 Data Hubs are a set of tools coupled with infrastructure that support four components: the submission, analysis, presentation and visualisation of SARS-CoV-2 raw read data, and its resulting analyses. What makes Data Hubs attractive is a unique set of features:
- Collaborative: Data Hubs enable data sharing amongst a group of collaborators in pre-publication, private mode, prior to publishing and data release.
- Configurable: Data Hubs enable researchers to use preferred tools for data submission, search and retrieval.
- Sustainable: Data Hubs are built on top of European Nucleotide Archive (ENA) infrastructure in order to reuse the same storage, data and metadata models.
A new publication in Microbial Genomics has been released describing the functionality of SARS-CoV-2 Data Hubs and specifically, a public SARS-CoV-2 Data Hub in detail.
The publication covers:
- Open data sharing through the European COVID-19 Data Platform
- Tools for submission, analysis, visualisation and data claiming (e.g. ORCiD)
- The systematic analysis of these datasets, at scale via the SARS-CoV-2 Data Hubs
- Lessons learnt
Using the COVID-19 Sequence Analysis Pipeline, the teams from Eotvos Lorand University (ELTE) in Hungary, Erasmus Medical Centre (EMC) in the Netherlands, Technical University of Denmark (DTU) and EMBL-EBI systematically analysed public raw read datasets generating consensus sequences and variant calls. These analysis products were then shared back to the public Data Hub where they are:
- visible through the COVID-19 Data Portal, enabling search and retrieval by the scientific community
- available for downstream databases and resources such as the European Variation Archive (EVA) and Ensembl
- available for downstream visualisation tools, such as the CoVEO Variant Browser.
What’s next?
Through the support of the VEO and BY-COVID projects, the SARS-CoV-2 Data Hubs are being expanded to support pandemic preparedness, using the data hubs foundations originally built under the COMPARE project. With this objective, the project team broadened their scope to cover more pathogens, which also implies more diseases, creating the new Pathogen Data Hubs, a component of the recently launched Pathogens Platform.
Based on the same concept containing four ‘mix and match’ components it is designed to support data sharing of pathogens beyond SARS-CoV-2. Visit the Pathogen Data Hubs dedicated page on the Pathogen Portal.
Acknowledgements
This work would not have been possible without:
- The extensive collaboration and contributions from the Eotvos Lorand University (ELTE) Hungary, Erasmus Medical Centre (EMC) Netherlands and Technical University of Denmark (DTU), along with EMBL-EBI.
- The large-scale open data sharing from various groups across the world, which have been highlighted through the data.
A full list of funders and partners can be found here.
Edit