We are pleased to announce the release of Ensembl 108, and the corresponding release of Ensembl Genomes 55 featuring changes in the human default tracks, new genomes in Ensembl Plants and Ensembl Metazoa, and the addition of mitochondrial annotation for Tasmanian devil.
Genome assemblies and annotation for many new species are also being continuously added to the Ensembl Rapid Release genome browser.
Default data display updates for human
In the Location tab, cDNAs, EST cluster (UniGene) and CCDS tracks will no longer be displayed by default. These tracks can still be added by configuring the page. See screen shots below.
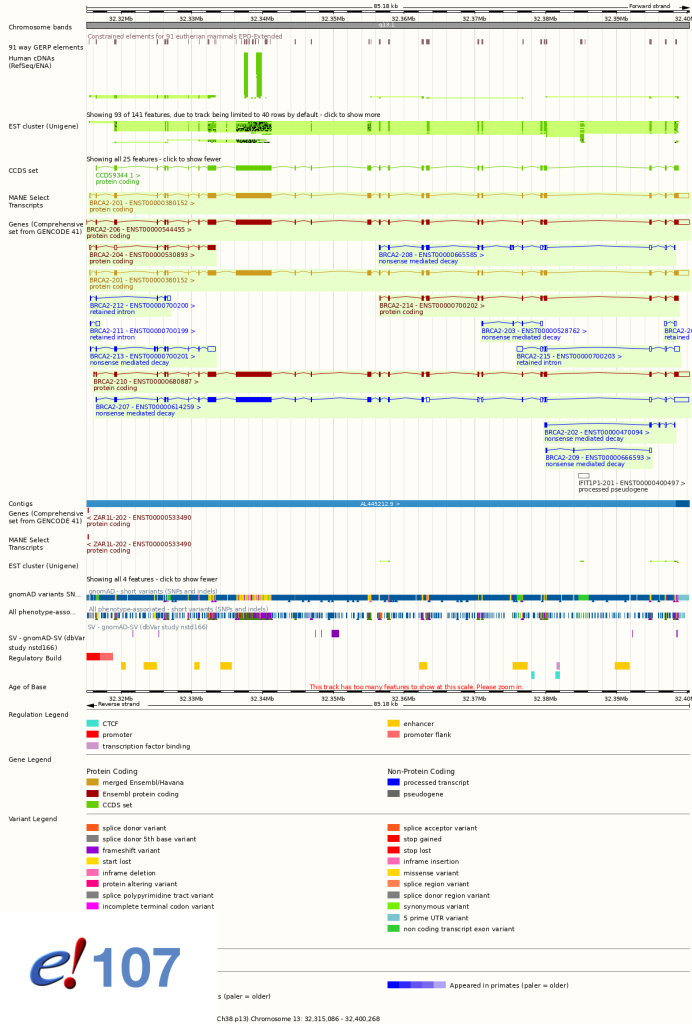
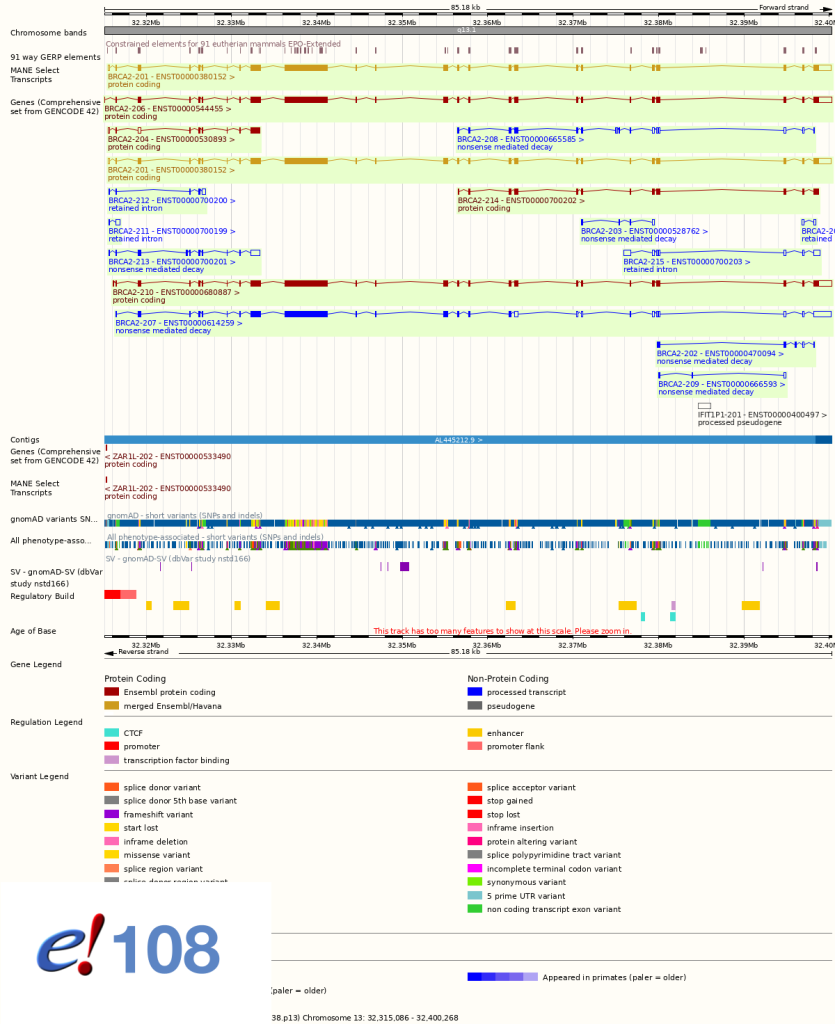
New genomes
Plants:
We have added two new species:
- Lolium perenne (Rye grass)
- Brassica juncea (Brown mustard)
Metazoa:
We have expanded the list of metazoa species available on Ensembl, which now includes:
- Amphibalanus amphitrite (Acorn barnacle)
- Acropora millepora (Stony coral)
- Acanthaster planci (Crown-of-thorns starfish)
- Actinia tenebrosa (Australian red waratah sea anemone)
- Asterias rubens (European starfish)
- Dimorphilus gyrociliatus (Meiobenthic segmented annelid)
- Eurytemora affinis (Calanoid copepod)
- Exaiptasia diaphana (Sea anemone)
- Homarus americanus (American lobster)
- Hyalella azteca (Amphipod)
- Hypsibius exemplaris (Water bear tardigrade)
- Mercenaria mercenaria (Hard clam)
- Mizuhopecten yessoensis (Yesso scallop)
- Octopus sinensis (East Asian common octopus)
- Owenia fusiformis (Polychaete annelid)
- Patiria miniata (Bat star starfish)
- Penaeus japonicus (Kuruma shrimp)
- Penaeus vannamei (Pacific white shrimp)
- Pollicipes pollicipes (Gooseneck barnacle)
- Portunus trituberculatus (Swimming crab)
- Procambarus clarkii (Red swamp crayfish)
New Assemblies and/or Annotation
Vertebrates:
- Mitochondrial annotation for Sarcophilus harrisii (Tasmanian devil, mSarHar1.11) have been added.
- RNASeq tracks including data from the GeneSWitCH consortium for Gallus gallus (chicken) reference (GRCg7b) and chicken breed (GRCg7w) have been added.
- Variation data from the European Variation Archive (EVA) are now displayed for five additional species: Macaca fascicularis (crab-eating macaque), Sander lucioperca (pike-perch), Microtus ochrogaster (prairie vole), Coturnix japonica (Japanese quail) and Ficedula albicollis (collared flycatcher).
Metazoa:
We have updated the assembly and genome annotation of:
- Daphnia magna (Fresh water planktonic crustacean)
- Lingula anatina (Lamp shell)
We have also updated the genome annotation of:
- Caenorhabditis elegans to WormBase annotation WS282.
- RefSeq variation annotation has been added to Octopus bimaculoides (California two-spot octopus)
Other updates and changes
- The Cyprinus carpio (Common carp) GCA_000951615.2 assembly has now been replaced with the newer Cypcar_WagV4.0 (GCA_905221575.1) assembly. The previous assembly has been retired from BioMart.
- Our variant databases for Bos taurus (cow), Danio rerio (zebrafish), Equus caballus (horse), Felis catus (cat), Gallus gallus (Red Jungle fowl chicken), Macaca mulatta (macaque), Mus musculus (mouse), Rattus norvegicus (rat) and Sus scrofa (pig) have been updated to use EVA release 3. This replaces previous data from dbSNP and uses new clustering and quality control methods, for example, variants which do not match the genome assembly will no longer be shown.
- MANE transcripts and proteins are now highlighted on gene and transcript summary pages. This gives more prominence to MANE and reduces the significance on CCDS.
- The transcribed_unprocessed_pseudogene biotype was changed to have separate pseudogene and lncRNA gene IDs in human and mouse, which increased the number of lncRNA genes in the total gene count.
- The biotype processed_transcript, when present in protein coding loci, has been replaced with protein_coding_CDS_not_defined.
- The Post-GWAS tool has been retired (read more here: https://www.ensembl.info/2022/07/28/retiring-the-post-gwas-tool/).
- Peak features have been removed from overlap REST endpoints, which may affect queries.
- Aphanomyces astaci, Aphanomyces invadans, Globisporangium ultimum, and Hyaloperonospora arabidopsidis have been removed from Ensembl Fungi, and fungal gene trees have been re-computed. These species will continue to be included in Ensembl Protists.
- Retirement of Ensembl 90 archive.